Large Cohort Transcriptomics
Adelaide Biomed Seminar Series
Black Ochre Data Labs
The Kids Research Institute Australia / ANU
March 13, 2026
Acknowledgement of Country

I would like to acknowledge that many of us are meeting today on Kaurna Country (Karrawirraparri: Redgum Forest River).
I acknowledge the deep feelings of attachment and relationship of the Kaurna people to their Place.
I also pay my respects to the cultural authority of Aboriginal and Torres Strait Islander peoples from other areas of Australia online today, and pay my respects to Elders past, present and emerging.
About Me
- Did 1st year science in 1986 \(\rightarrow\) completed 2002-2005 (Genetics)
- BMa.Comp.Sc. (Hons) 2006-2007
- 1988-1991: Elder Conservatorium
- First came across R (1.5.1) analysing two-colour microarrays in Dec 2002

About Me
- PhD 2008-2018
- Normal student: 2008-2011
- Homeless/Couch-surfing: 2012-2013
- Bioinformatics Hub: 2014-2020
- 2020-2022: Dame Roma Mitchell Cancer Research Labs
- 2022-2026: Black Ochre Data Labs
Bioconductor Enthusiast


![]()


- Also developed & maintain
strandcheckR(Hien To)- Helped with
sSNAPPY(Nora Liu) +tadar(Lachlan Baer)
- Helped with
- Currently co-chair of Community Advisory Board
- Leading BiocAsia Working Group


The PROPHECY Study
The PROPHECY Study
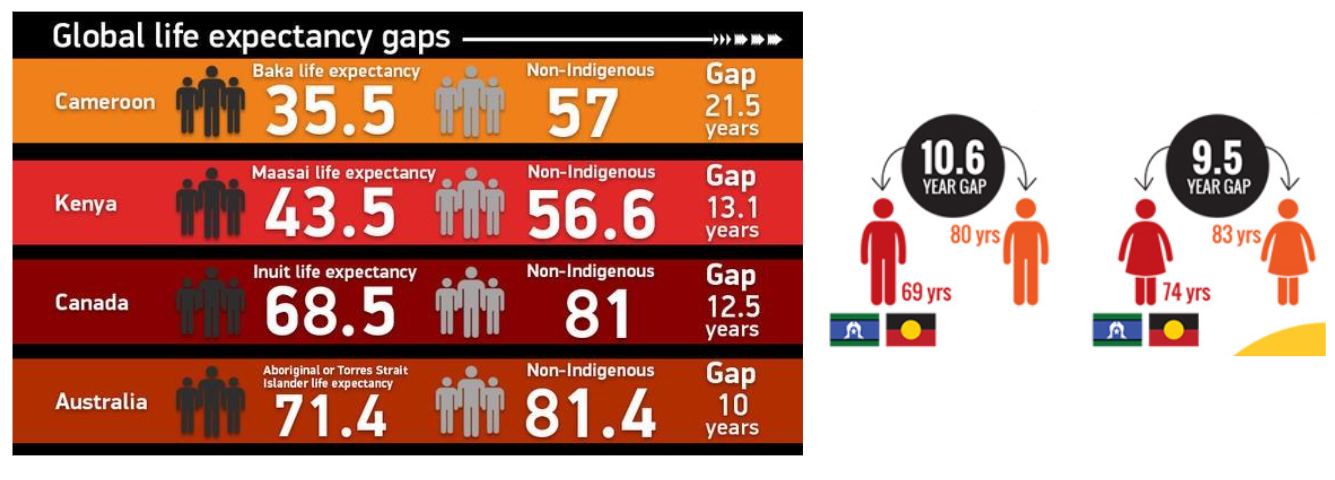
The PROPHECY Study
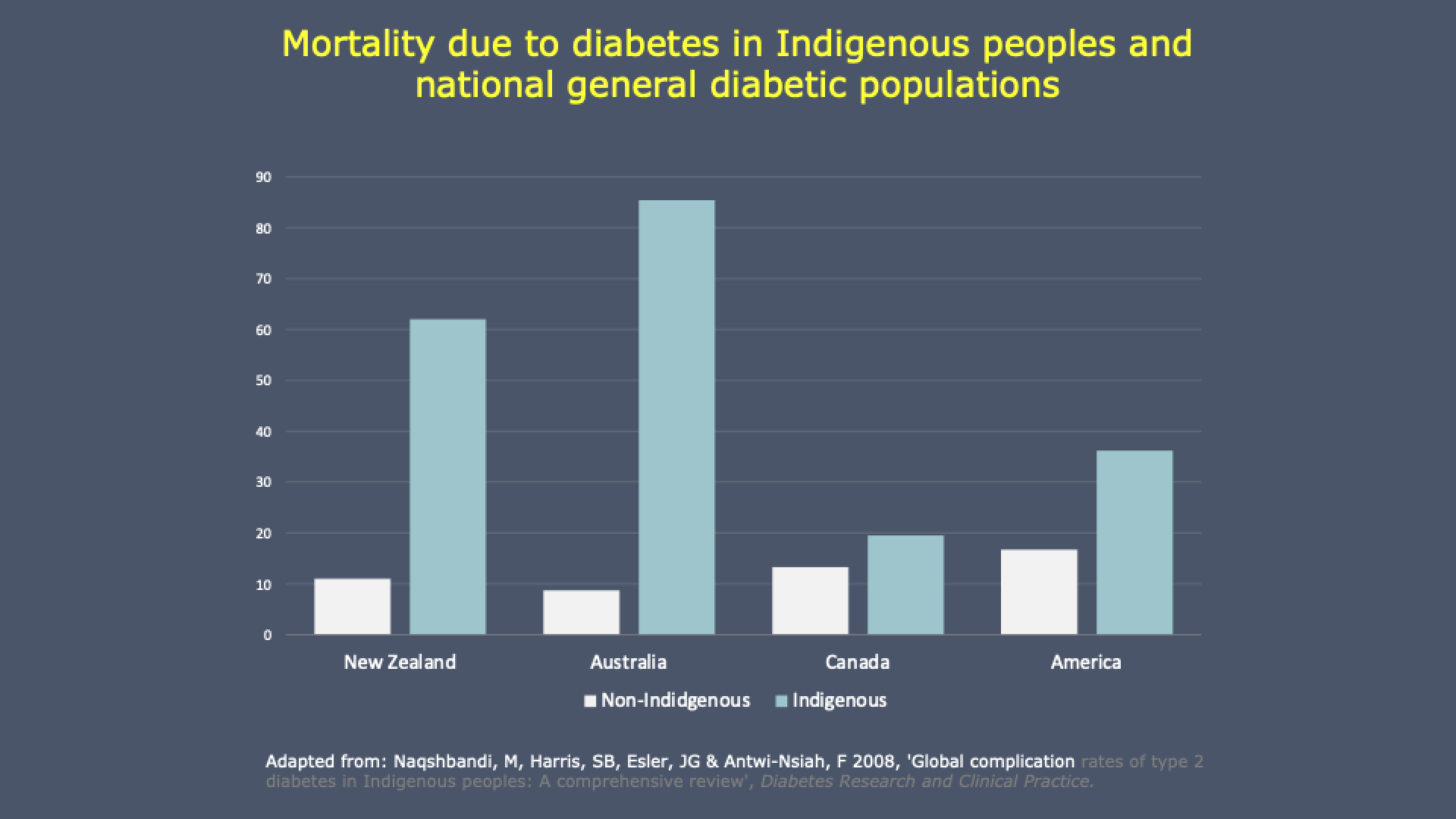
The PROPHECY Study
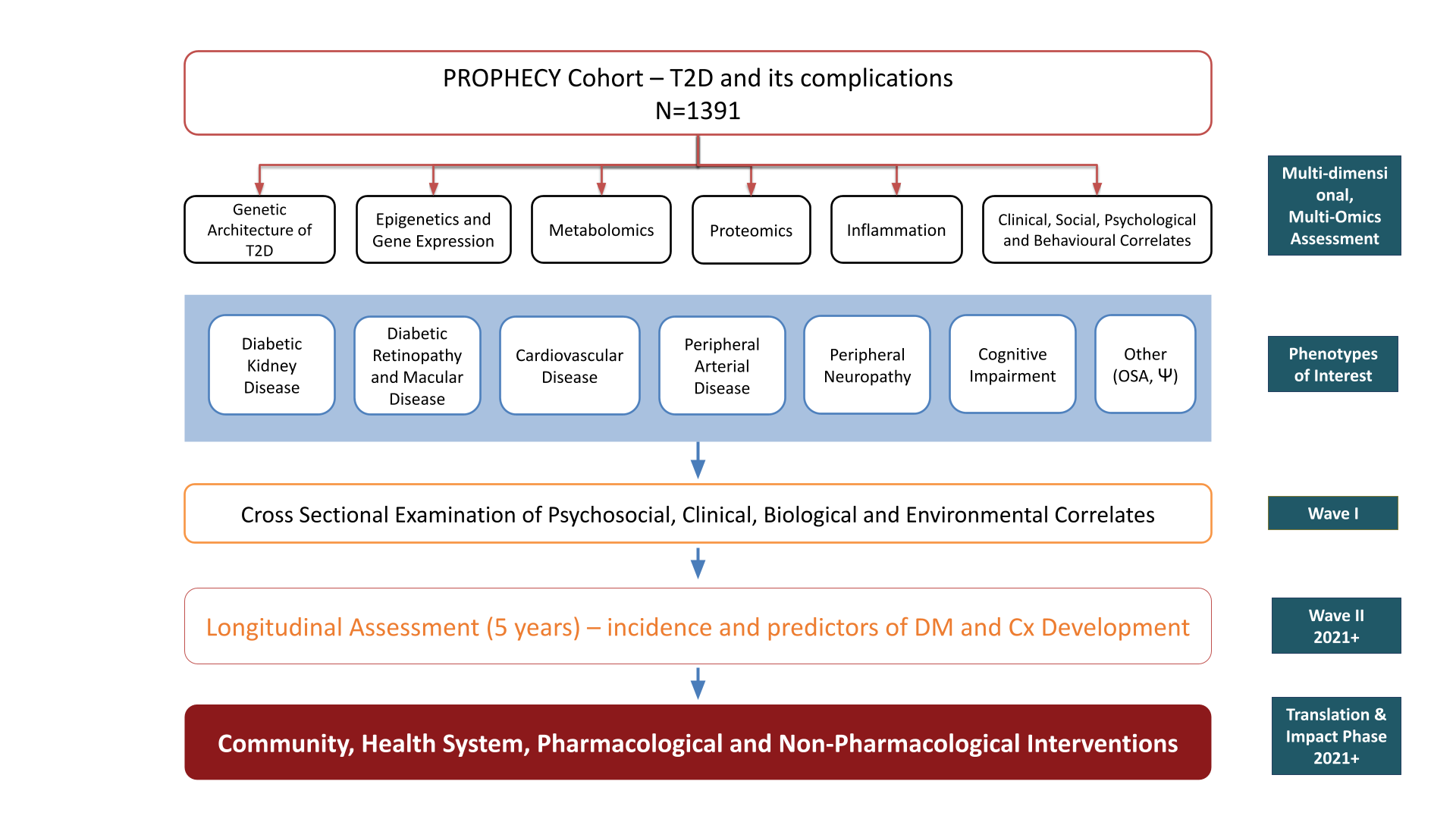
The PROPHECY Study
- A key design element \(\rightarrow\) all research is community led, not researcher led
- Prof Alex Brown spent 5 years consulting with SA communities first
- Strong Indigenous Data Sovereignty Principles
- Strict data management protocols
- High demand for clear documentation

Clickbait Time
Is large cohort transcriptomics just scRNA without the zero counts?

The PROPHECY Transcriptomics Layer
Sequencing Considerations
- Whole Blood from participants \(\implies\) incredibly valuable resource
- Trying to to get everything as ‘right’ as possible \(\implies\) no second chances
- 13x96 well plates were planned for & paid before I joined \(\implies\) N = 1248

- Subset of 93 participants already sequenced by the El Osta group
- Focus on Chronic Kidney Disease (CKD) + Diabetes (T2D)
- Total RNA \(\implies\) rRNA Depleted but not hbRNA depleted
- hbRNA was 30-50% of library
- rRNA consistently below 20% (mostly below 10%)
- Pilot study compared polyA with totalRNA (rRNA + hbRNA depleted)
Study Design
- Prepared in batches of 96 \(\implies\) batch effects
- All reagents from same supplier batches
- Chose a design strategy compatible with hRUV (Kim et al. 2021)
- Short replicates, Long replicates, Pools
- 1134 Blood samples were selected for inclusion based on:
- RIN >5.0 & RNA concentration > 25ng/μl
- Samples are stored in extraction order with a ULN
- Randomising samples would be extremely difficult
- Only had access to RIN, Concentration & Sex
- El Osta sub-cohort were all late ULNs
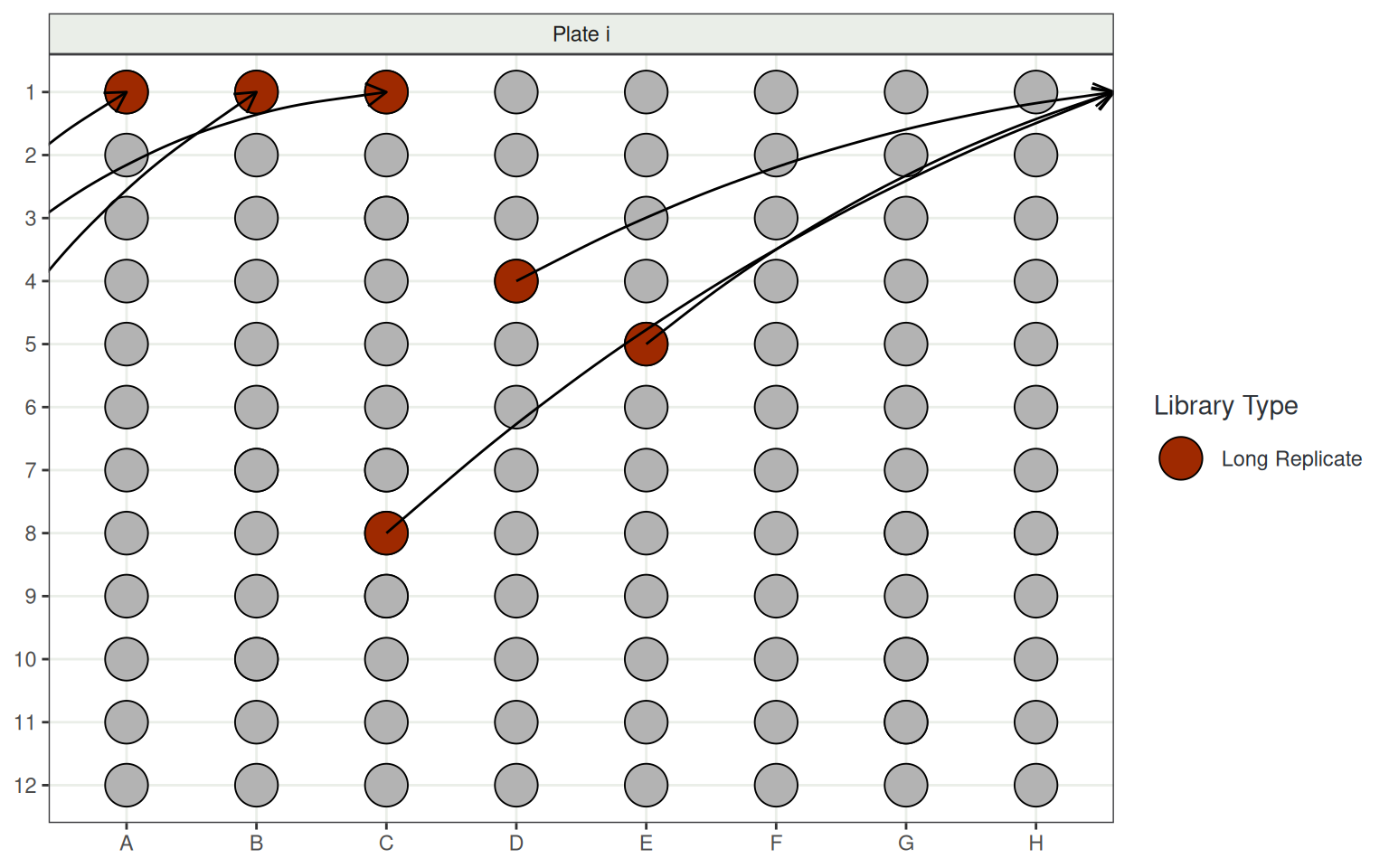
Experimental Design
- Plates include:
- 6 Long Replicates to straddle consecutive plates
Trivia Time!!!
- Did you know that plates are prepared using a multi-tip pipette?
- Ordered by columns not rows
Experimental Design
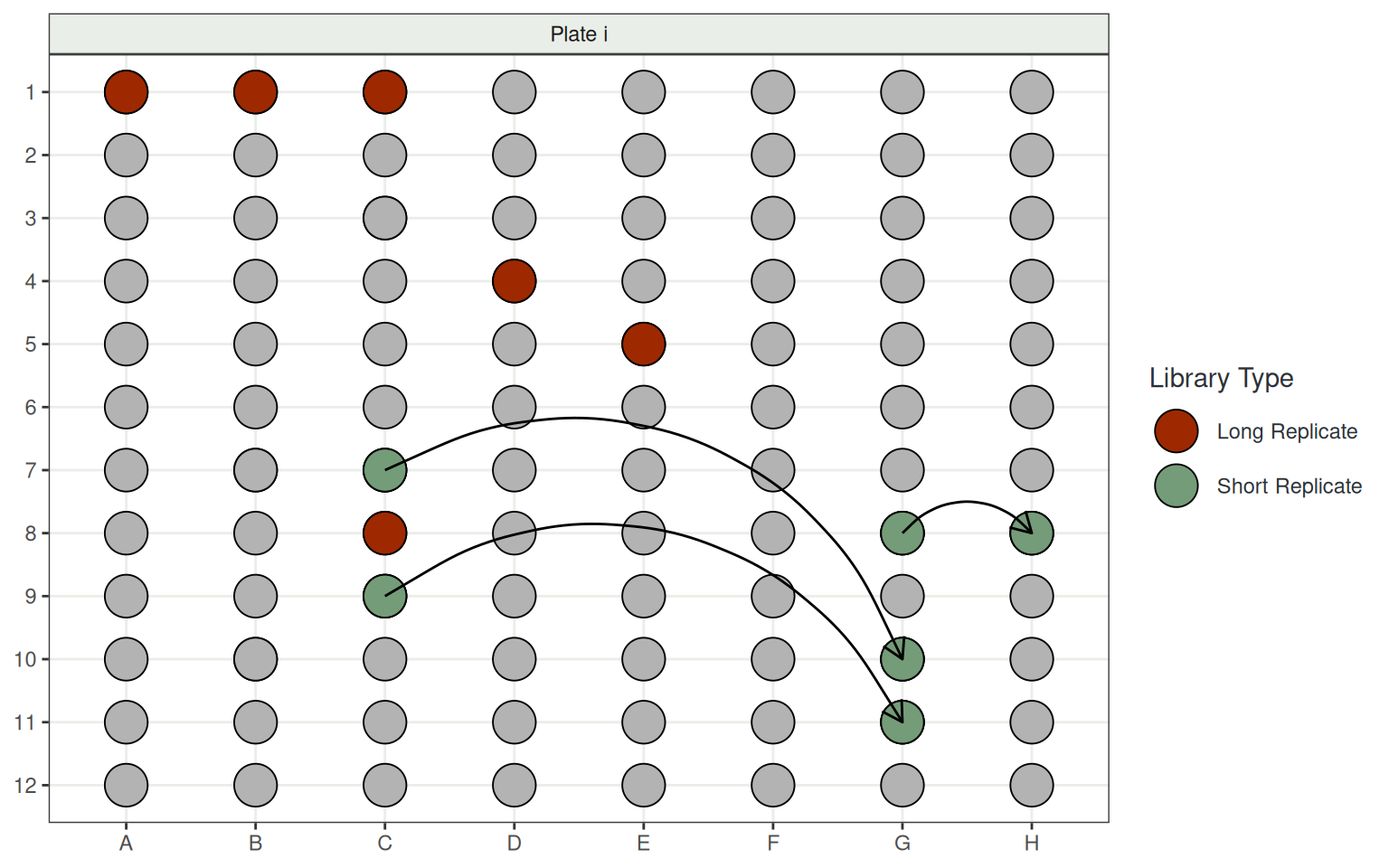
- Plates include:
- 6 Long Replicates to straddle consecutive plates
- 6 Short Replicates within plates
Experimental Design
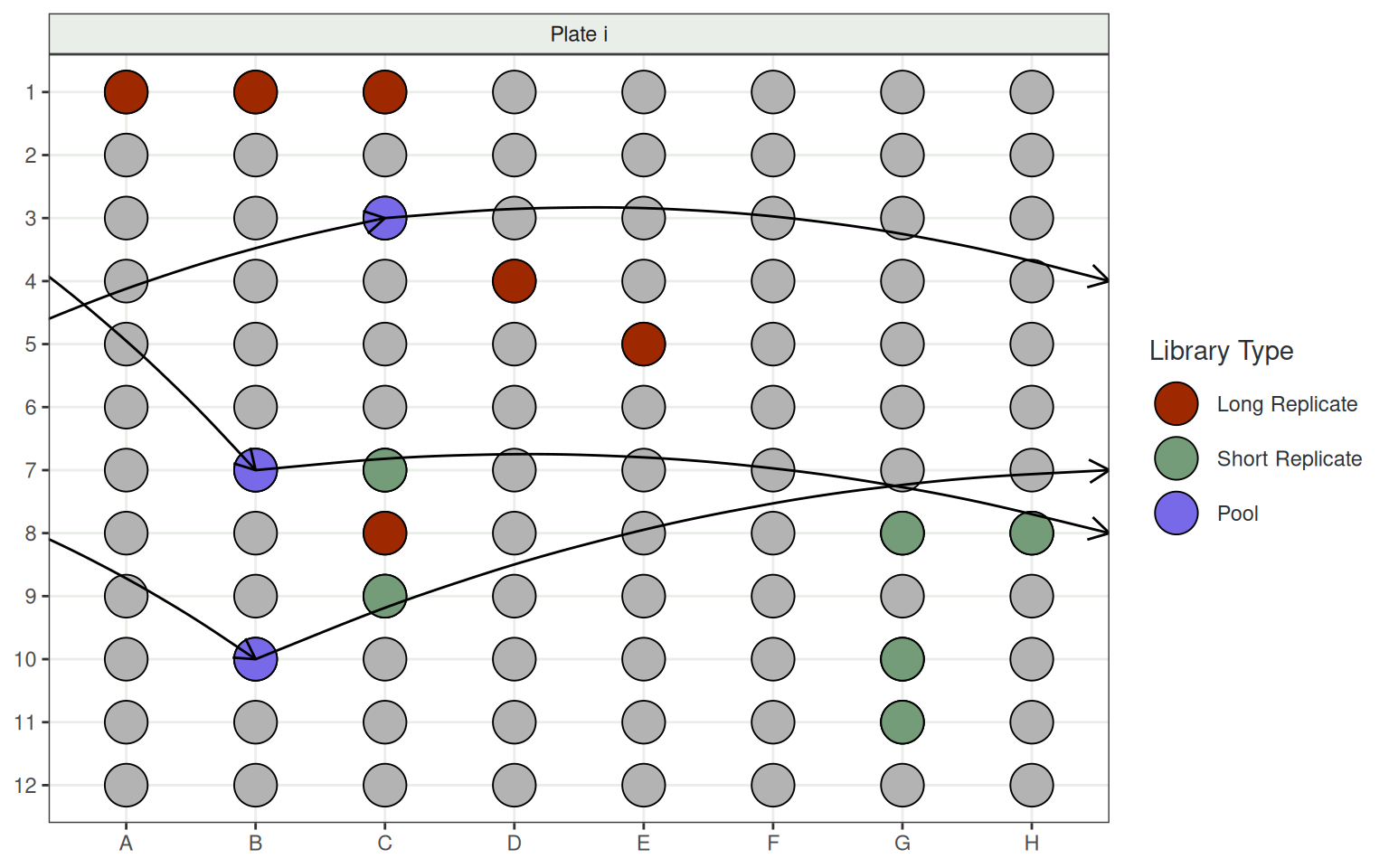
- Plates include:
- 6 Long Replicates to straddle consecutive plates
- 6 Short Replicates within plates
- 3 distinct pools across all 13 plates
- 87 Participants \(\implies\) 1,134 total
Data Processing Workflow
- Data processing written in
nextflow- Dr Alastair Ludington & Holly Massacci
- Run as every plate came in
- Produced a set of plate-level QC reports
- Threw everything at it:
fastp,RSeQC,samtools stats, all log files etc - Forgot to include
strandcheckR(To and Pederson 2019) \(\rightarrow\) supplementary pipeline
- Threw everything at it:
- Manually checked every plate for Pass/Fail/Caution
- Thinking about which libraries to resequence


Data Processing Workflow
Pipeline Output
/path/to/prophecy/PROP000000/ # PROPHECY ID
└── 1234567 # Aliquot ID
└── RNASEQ
├── bigwigs
│ └── E200099999 # Sequencing Run ID
│ └── 23-00000 # ULN
│ ├── GRCh38
│ └── PROPHECY
├── depletion
│ └── E200099999
│ └── 23-00000
│ ├── 1234567_1_hbrna.flagstat
│ └── 1234567_1_rrna.flagstat
├── fastp
│ └── E200099999
│ └── 23-00000
│ ├── 1234567_1_fastp.html
│ └── 1234567_1_fastp.json
├── fastq
│ └── E200099999
│ └── 23-00000
│ ├── 23-00000_1_S24_L01_R1_001.fastq.gz
│ ├── 23-00000_1_S24_L01_R1_001.md5
│ └── 23-00000_1_S24_L01_R2_001.fastq.gz
├── featureCounts
│ └── E200099999
│ └── 23-00000
│ ├── GRCh38
│ └── PROPHECY
├── rseqc
│ └── E200099999
│ └── 23-00000
│ ├── GRCh38
│ │ ├── inner_distance
│ │ │ ├── 1234567_1.inner_distance_freq.txt
│ │ │ ├── 1234567_1.inner_distance_mean.txt
│ │ │ └── 1234567_1.inner_distance.txt
│ │ ├── read_distribution
│ │ │ └── 1234567_1.read_distribution.tsv
│ │ └── tin
│ │ ├── 1234567_1.summary.txt
│ │ └── 1234567_1.tin.tsv
│ └── PROPHECY
├── salmon
│ └── E200099999
│ └── 23-00000
│ ├── GRCh38
│ │ └── 1234567_1
│ │ ├── aux_info
│ │ ├── cmd_info.json
│ │ ├── lib_format_counts.json
│ │ ├── libParams
│ │ ├── logs
│ │ └── quant.sf
│ └── PROPHECY
└── star
└── E200099999
└── 23-00000
├── GRCh38
│ ├── 1234567_1.cram
│ ├── 1234567_1.cram.crai
│ ├── 1234567_1.cram.md5
│ ├── 1234567_1.Log.final.out
│ ├── 1234567_1.SJ.out.tab
│ ├── 1234567_1.stats
│ └── 1234567_1_strandcheckr.tsv.gz
└── PROPHECY- For all 1248 libraries (including 1 repeated run …)
Final Outcome
- 31 sample mixups \(\implies\) all resolved successfully
- Only one library was an unexpected participant
- 17 had no genotypes \(\implies\) assumed correct
- 1 library broke
STARconsensusDUGteam identified bug inSTAR& made pull request
- Re-integrating mixups was non-trivial
- QC reports written for 96 samples failed
- Manual re-collation of all QC outputs
Final Summary
| Plate | Pass | Caution | Fail | Incomplete | Total Yield (billions) | Mean Q30 |
|---|---|---|---|---|---|---|
| SAGCQA0503-2 | 40 | 47 | 9 | 0 | 6.26 | 94.8% |
| SAGCQA0503-3 | 92 | 4 | 0 | 0 | 6.36 | 95.2% |
| SAGCQA0503-4 | 70 | 15 | 11 | 0 | 6.14 | 94.4% |
| SAGCQA0503-5 | 68 | 18 | 10 | 0 | 6.24 | 94.2% |
| SAGCQA0503-6 | 62 | 28 | 6 | 0 | 6.31 | 94.7% |
| SAGCQA0503-7 | 77 | 11 | 8 | 0 | 5.94 | 94.2% |
| SAGCQA0503-8 | 85 | 6 | 5 | 0 | 5.51 | 92.4% |
| SAGCQA0503-9 | 92 | 1 | 3 | 0 | 5.81 | 93.1% |
| SAGCQA0503-10 | 68 | 22 | 6 | 0 | 5.69 | 93.3% |
| SAGCQA0503-11 | 88 | 7 | 0 | 1 | 4.89 | 91.4% |
| SAGCQA0503-12 | 49 | 36 | 11 | 0 | 5.83 | 93.9% |
| SAGCQA0503-13 | 90 | 3 | 3 | 0 | 5.21 | 92.8% |
| SAGCQA0503-14 | 38 | 52 | 6 | 0 | 6.31 | 94.7% |
| Total | 919 | 250 | 78 | 1 |
QC
| Name | Source | Interpretation |
|---|---|---|
status |
Manual Inspection | Originally defined in order to identify libraries which might be candidates for resequencing |
rin |
SAGC | RNA Integrity Number with high values (> 7) indicating a higher quality sample |
conc |
SAGC | RNA Concentration in the original extractions |
depletion_conc |
SAGC | RNA Concentration used for the rRNA/hbRNA deletion step. Two values were used with the lower concentration being an attempt to reduce saturation of the depletion reagents |
rrna_rate |
BODL pipeline | Estimates the proportion of reads deriving from rRNA by aligning a subset of reads to an rRNA reference |
hbrna_rate |
BODL pipeline | Estimates the proportion of reads deriving from hbRNA by aligning a subset of reads to an rRNA reference |
duplication_rate |
fastp |
Taken directly from the fastp reports, given that UMIs were included in all libraries |
sequenced_fragments |
fastp |
The initial number of fragments contained in each pair of fastq files |
q30_rate |
fastp |
The proportion of bases acheiving a PHRED quality score > 30 when being sequenced |
gc_content |
fastp |
The average GC content of reads across the entire library |
gene_assigned_rate |
salmon |
num_frags_with_concordant_consistent_mappings / sequenced_fragments |
decoy_aligned_rate |
salmon |
num_decoy_fragments / sequenced_fragments |
median_insert_size |
samtools stats |
Calculated by only taking alignments which are concordantly paired |
sd_log10_insert_size |
samtools stats |
Calculated by only taking alignments which are concordantly paired, the transforming the insert size using log10 and finding the standard deviation |
median_tin |
nf-core |
The median TIN score across all measured trancripts |
prop_no_feature |
featureCounts |
The proportion of input reads which aligned where there was no annotated feature |
stranded_coverage |
strandcheckR |
The proportion of windows showing >90% of reads with a stranded bias, weighted by coverage |
prop_assigned |
featureCounts / salmon |
The proportion of sequenced fragments assigned to a gene/transcript |
QC: Duplication Rates
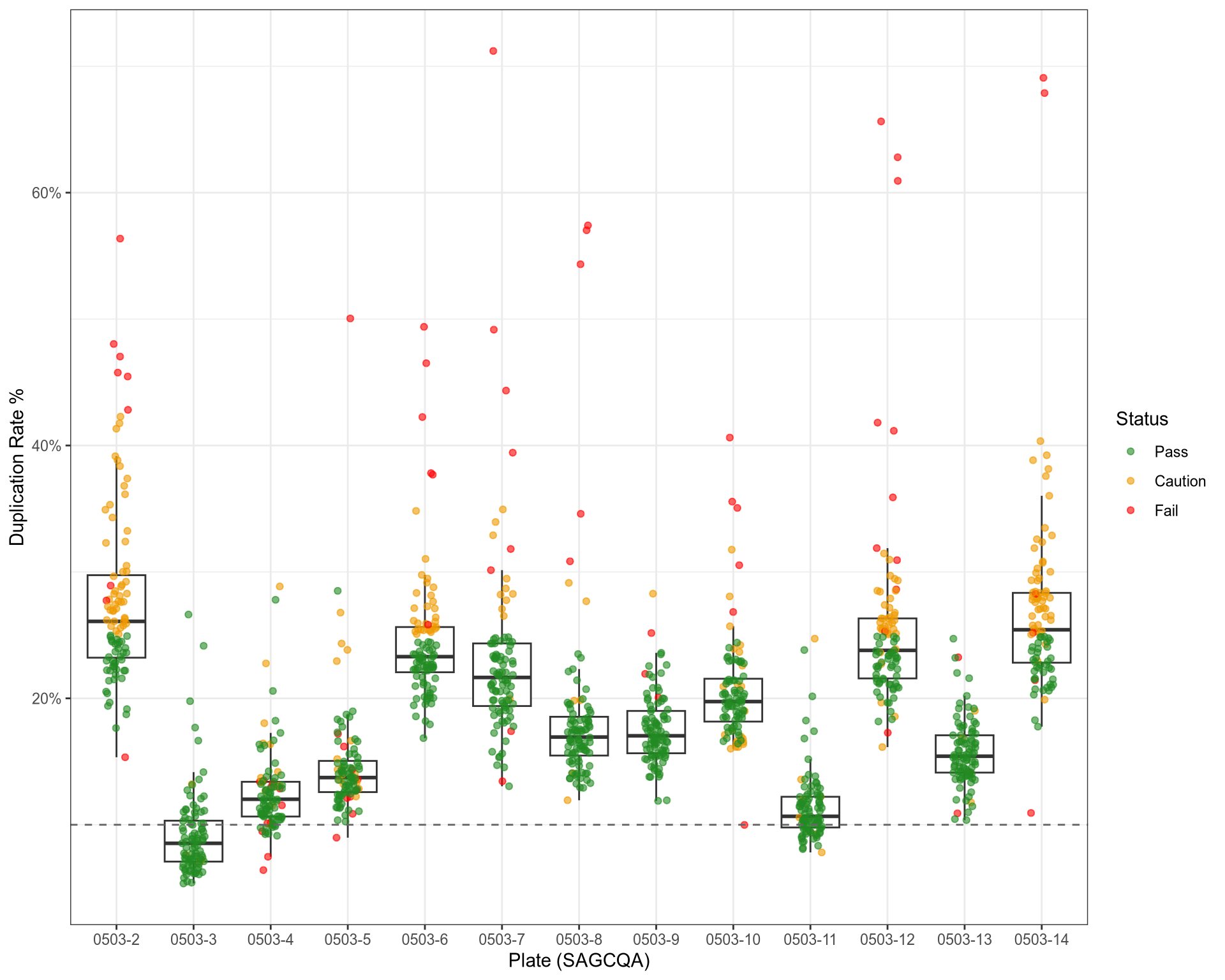
QC: rRNA Content
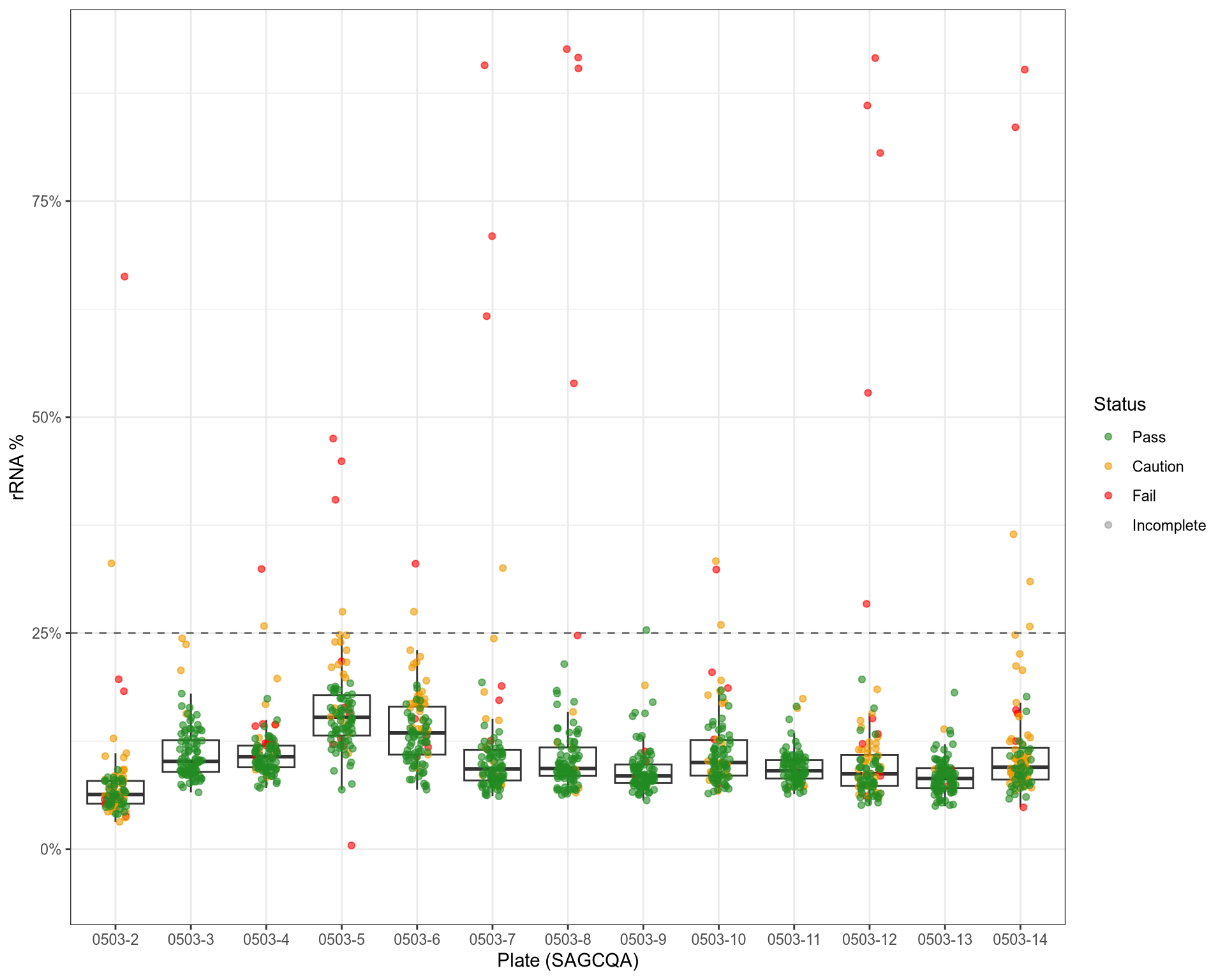
QC: Strandedness of Alignments
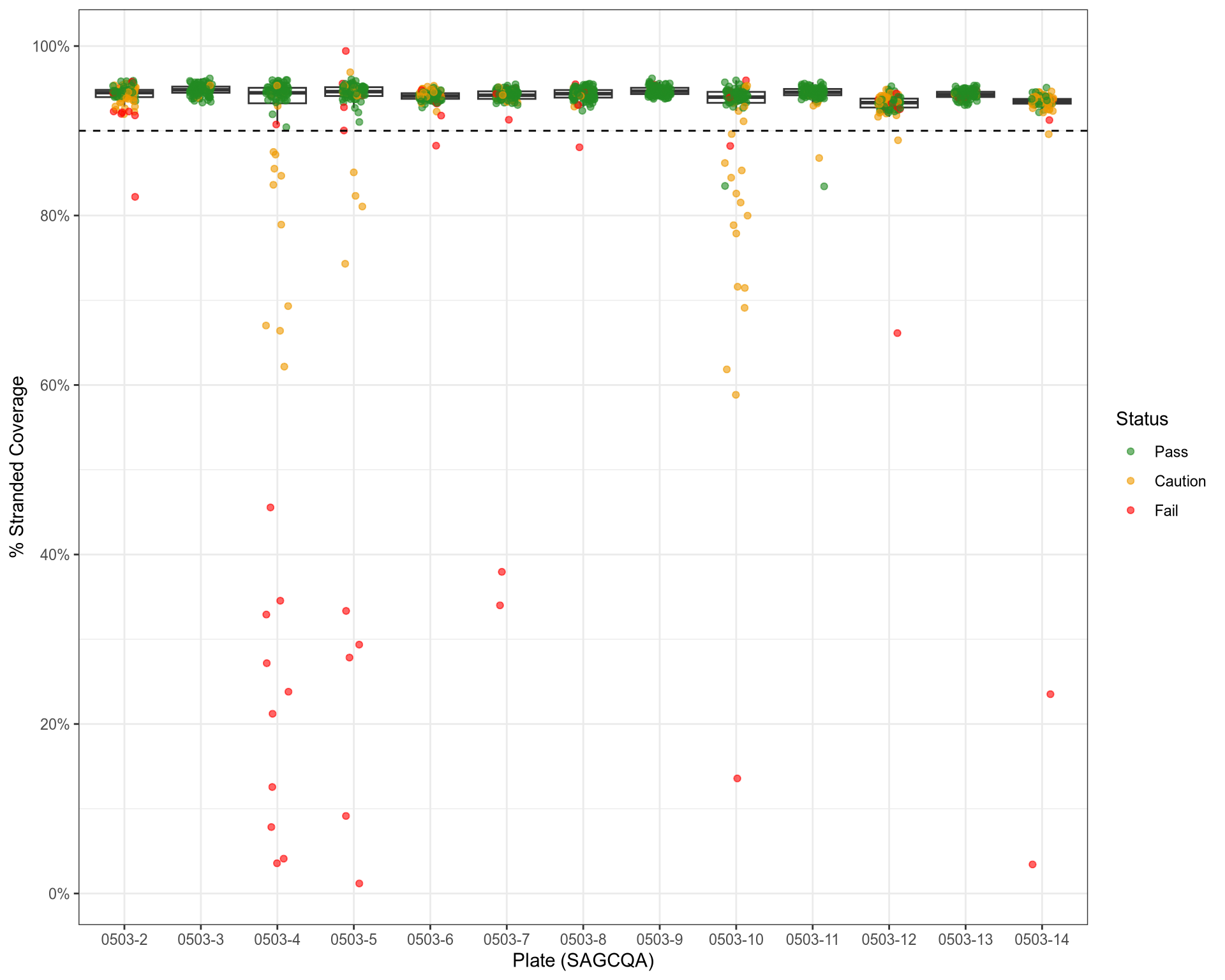
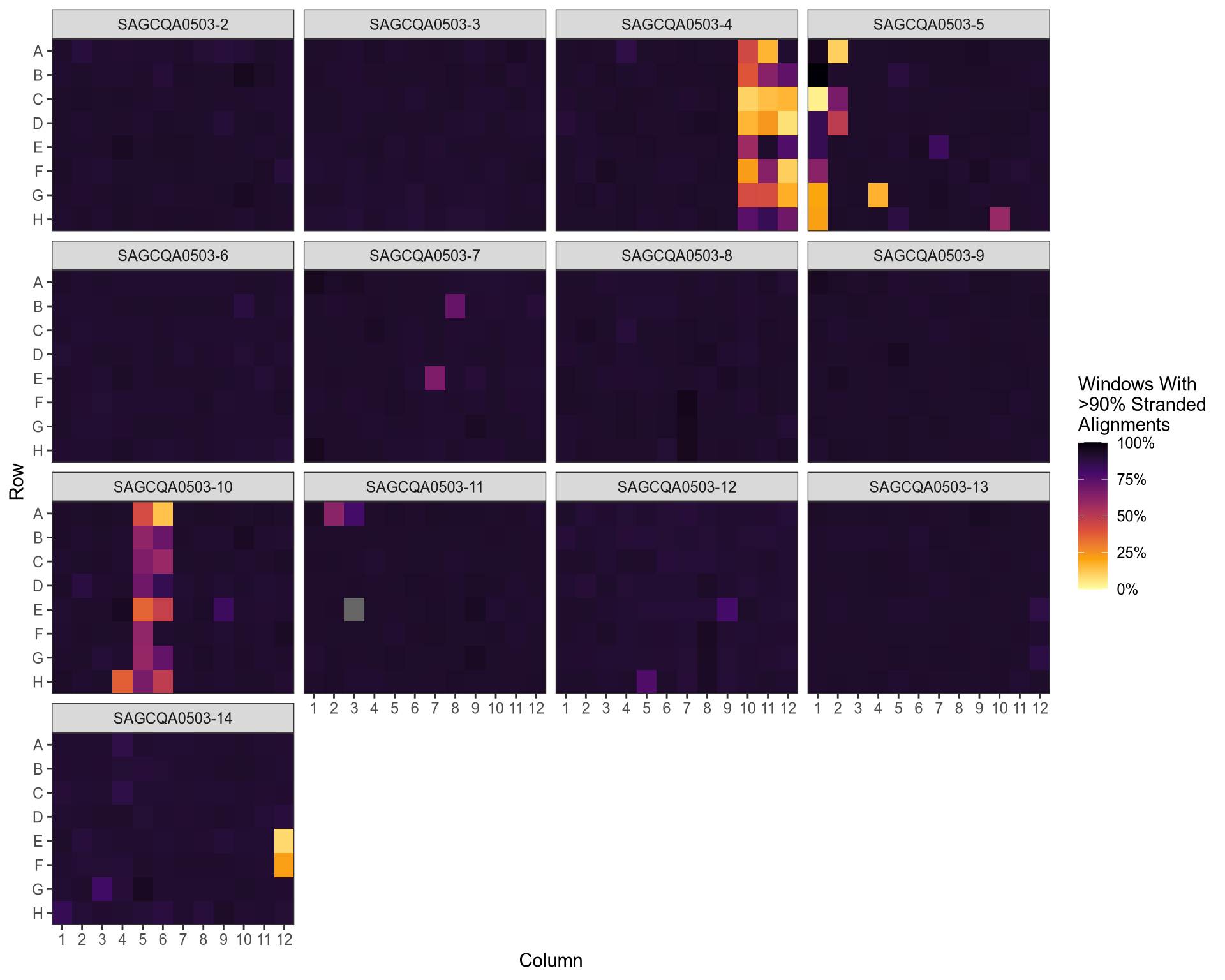
QC: Strandedness Vs Insert Size Variability
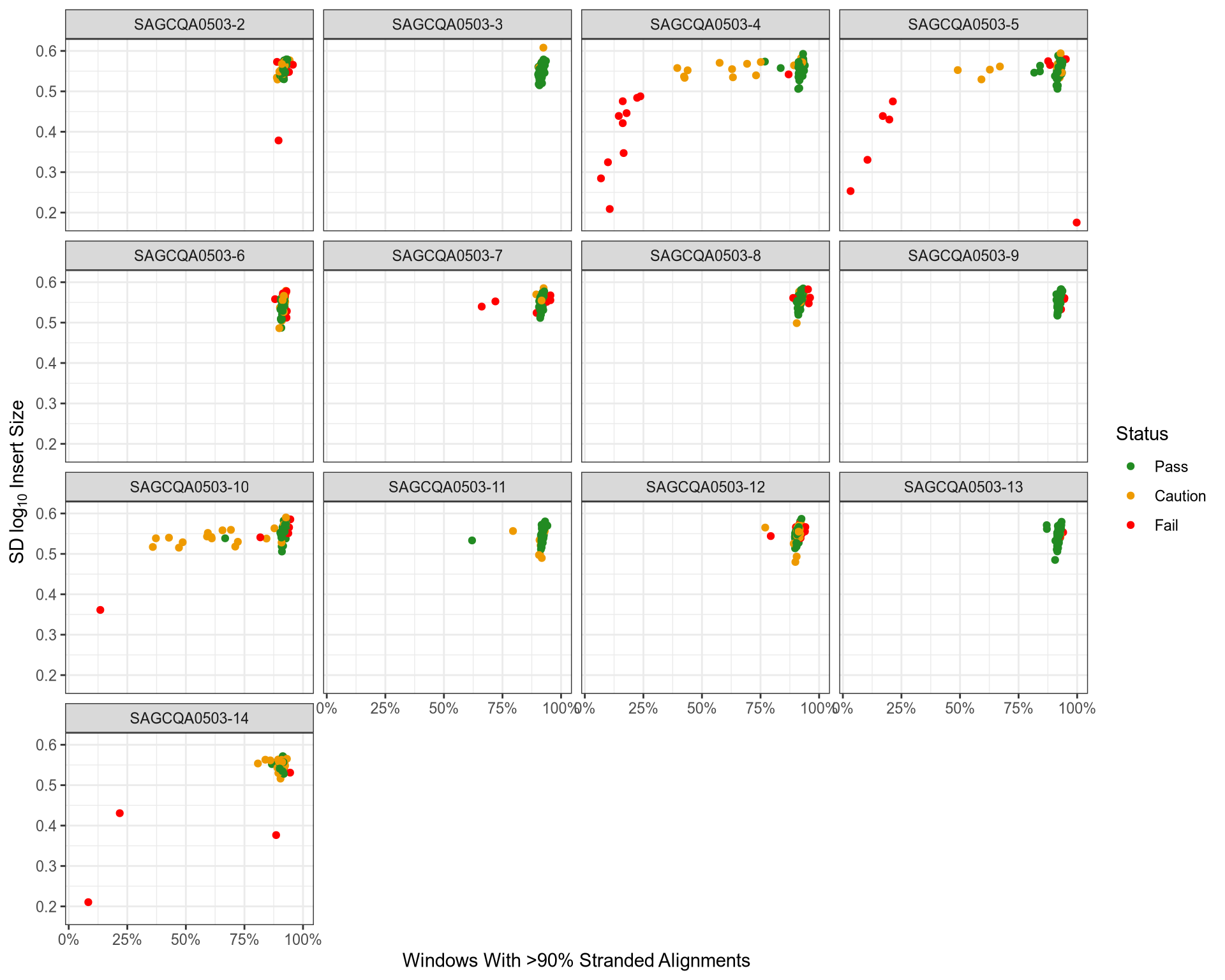
QC: % Of Sequenced Reads Assigned to Genes
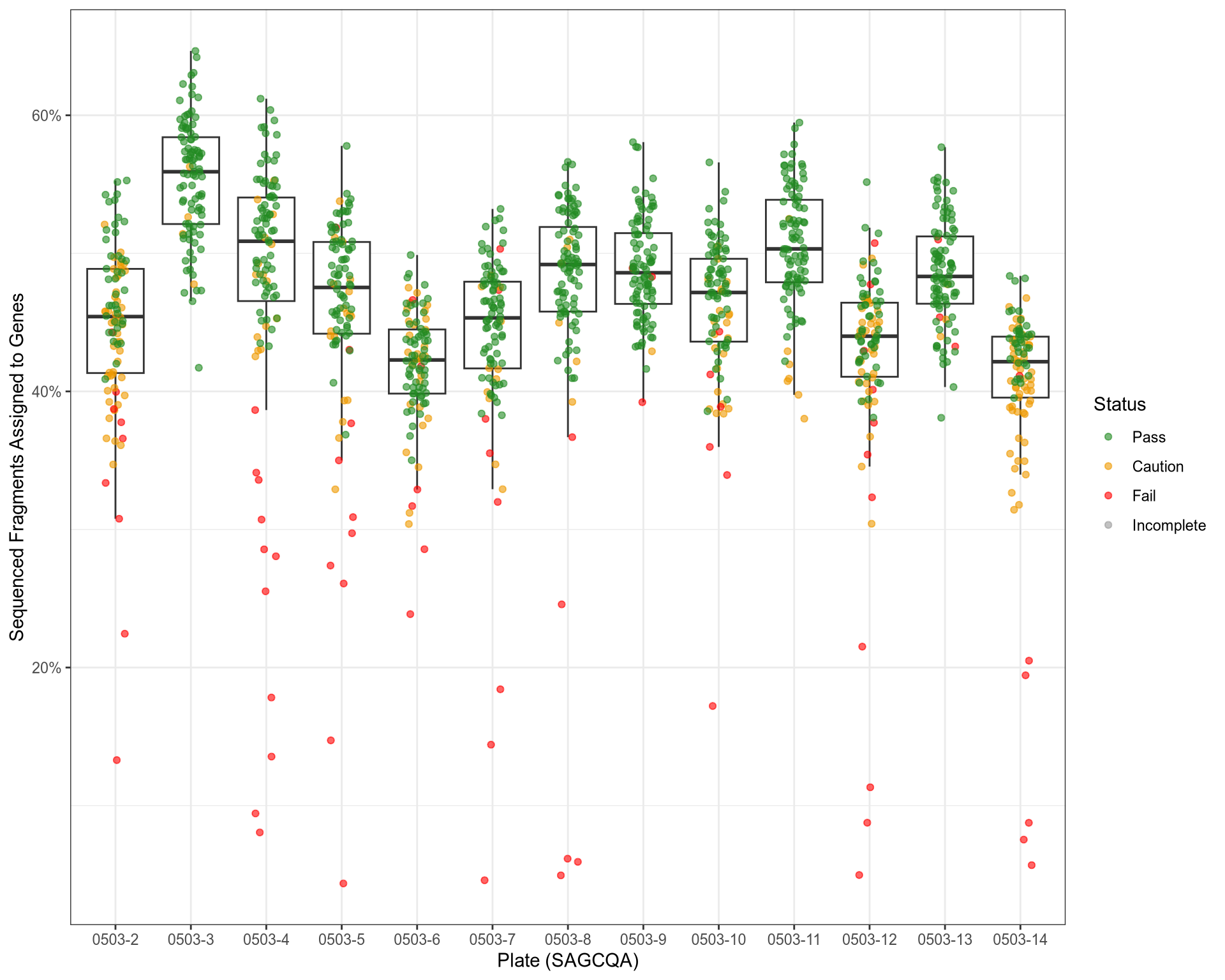
Similarity of QC Metrics
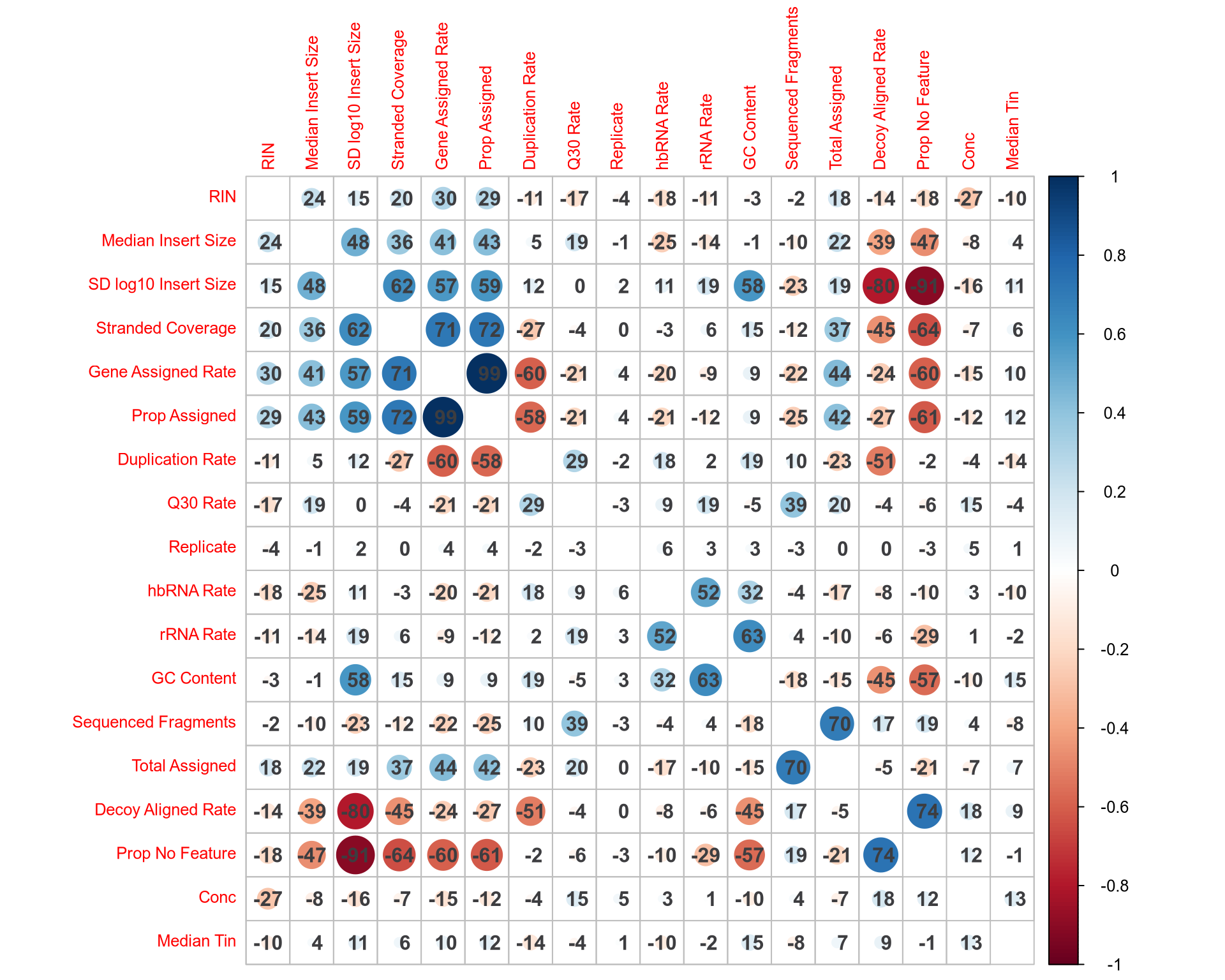
Outlier Detection
- Run PCA
- Find Robust Z-scores for PC1-4
- Label as “No”, “Potential” or “Unambiguous” Outlier
- |Z| > 1/200 (2.5758) or |Z| > 1/2000 (3.2905)
- Find which QC metrics have outlier threshold
Outlier Detection
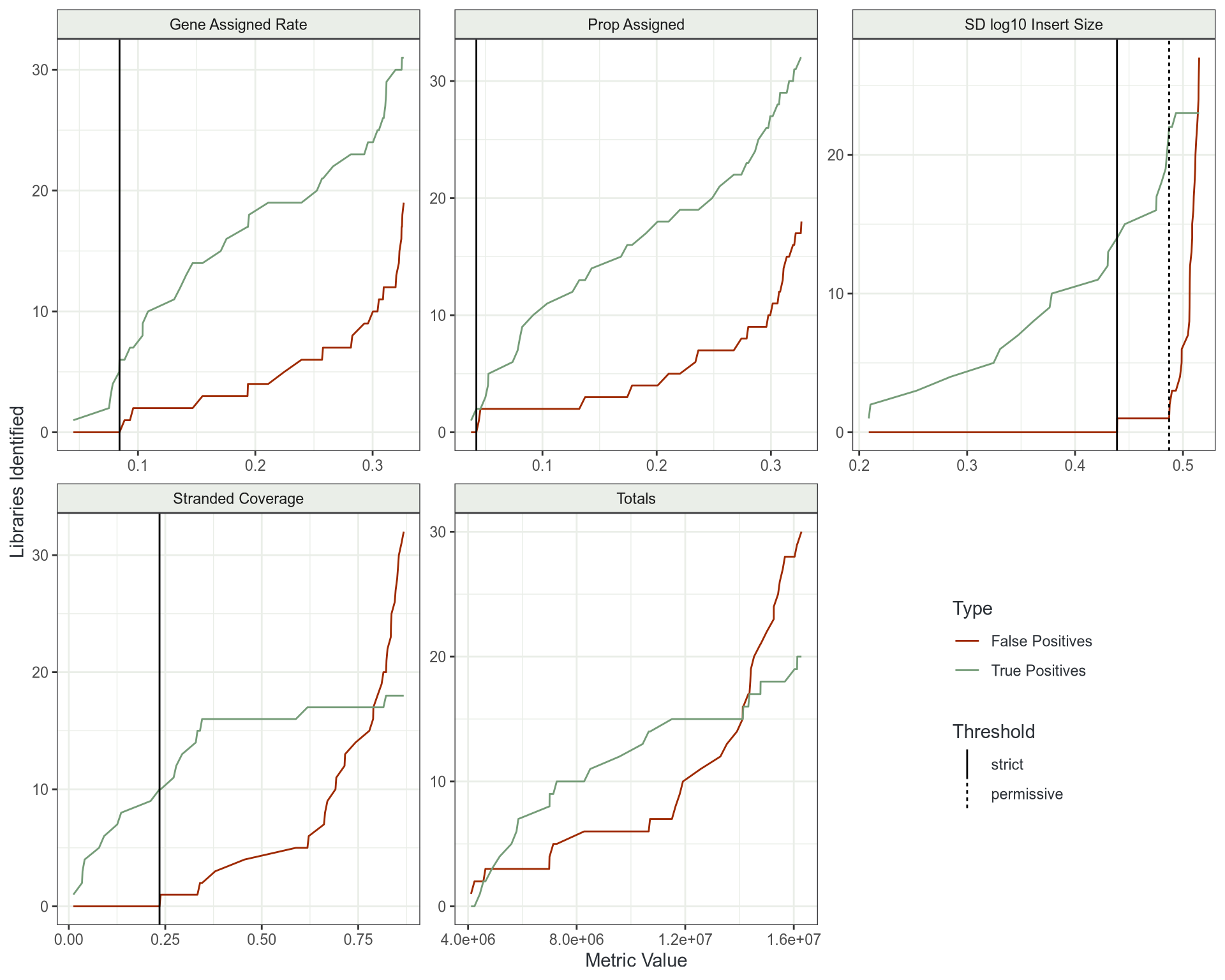
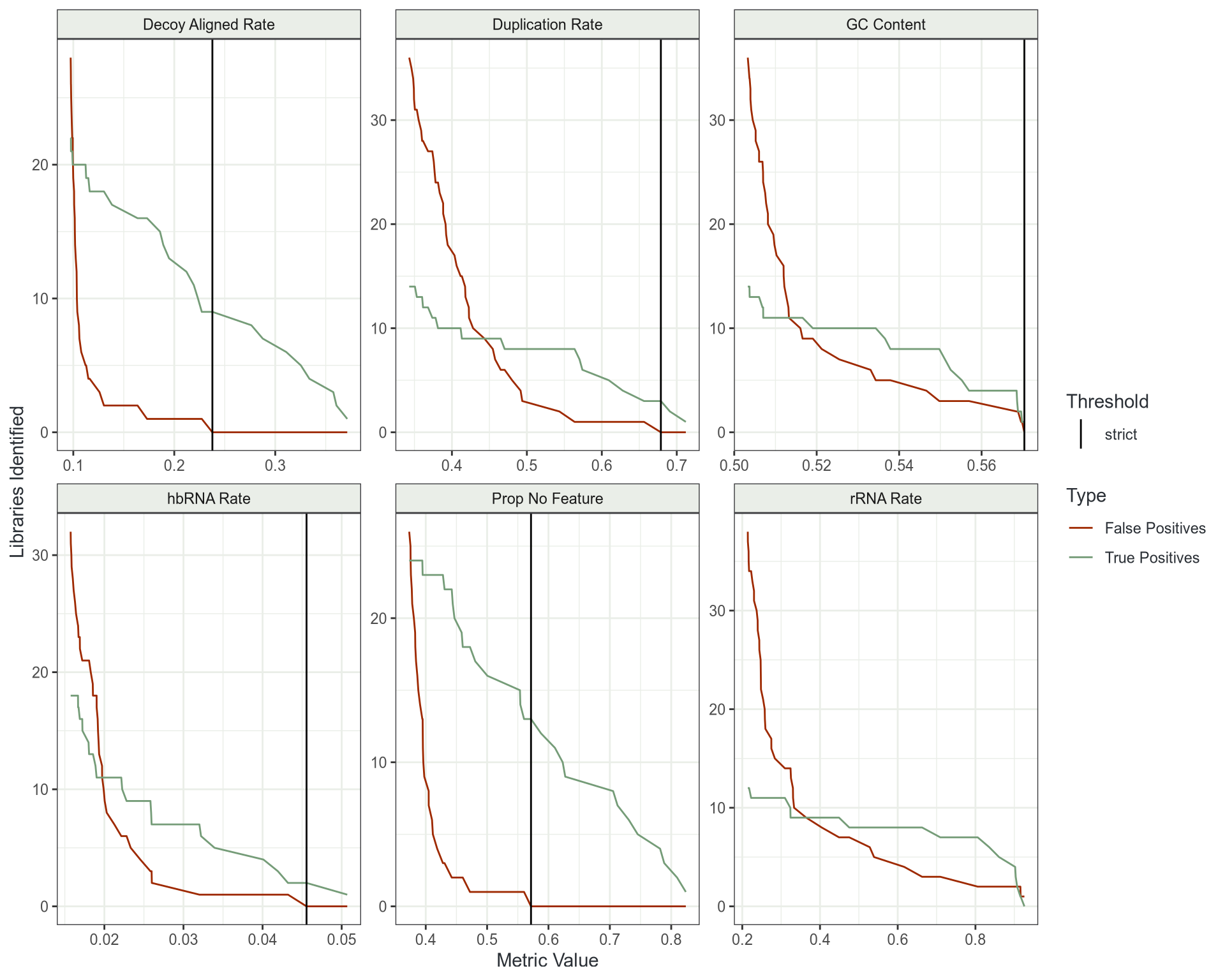
Outlier Detection
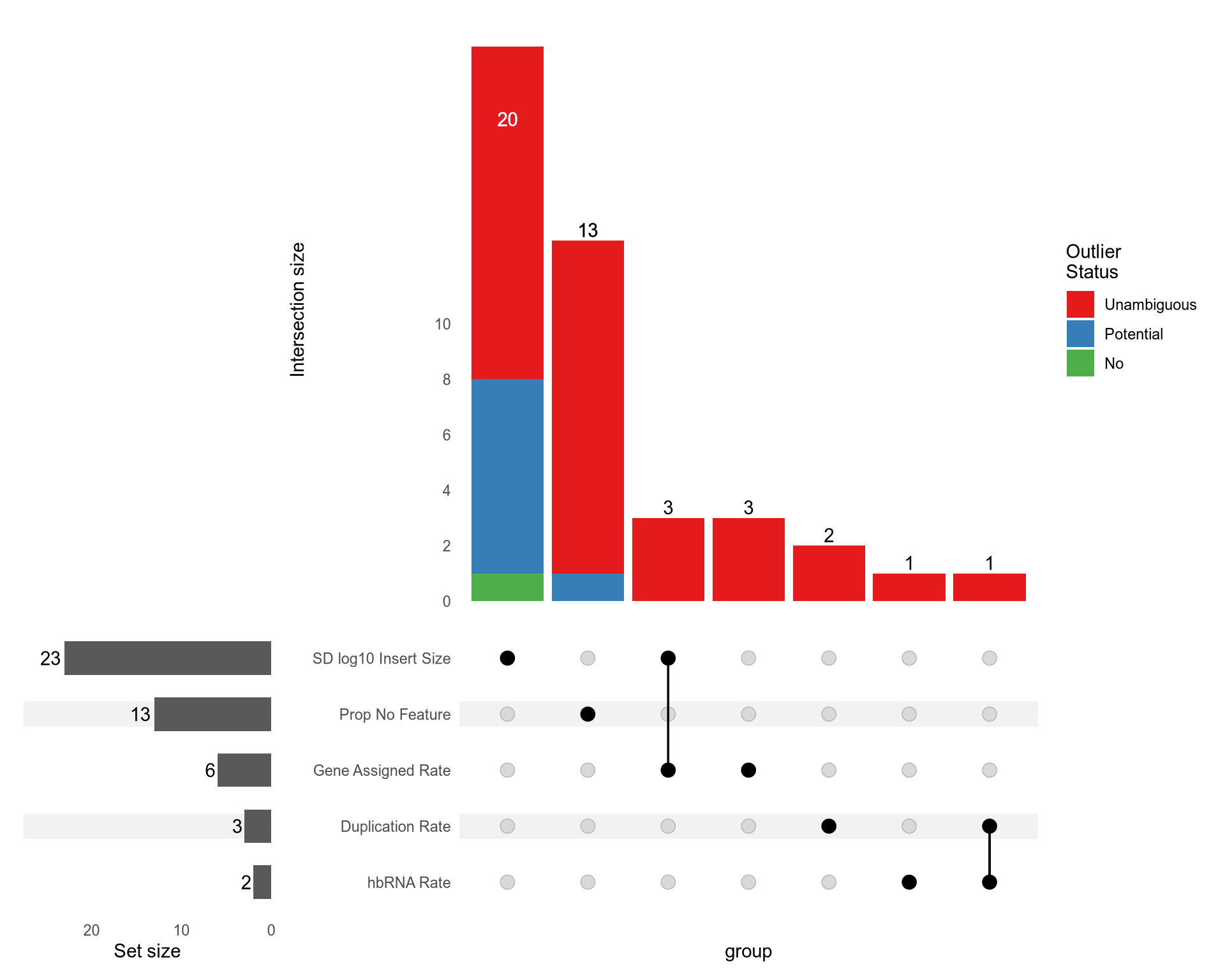

Outlier Detection
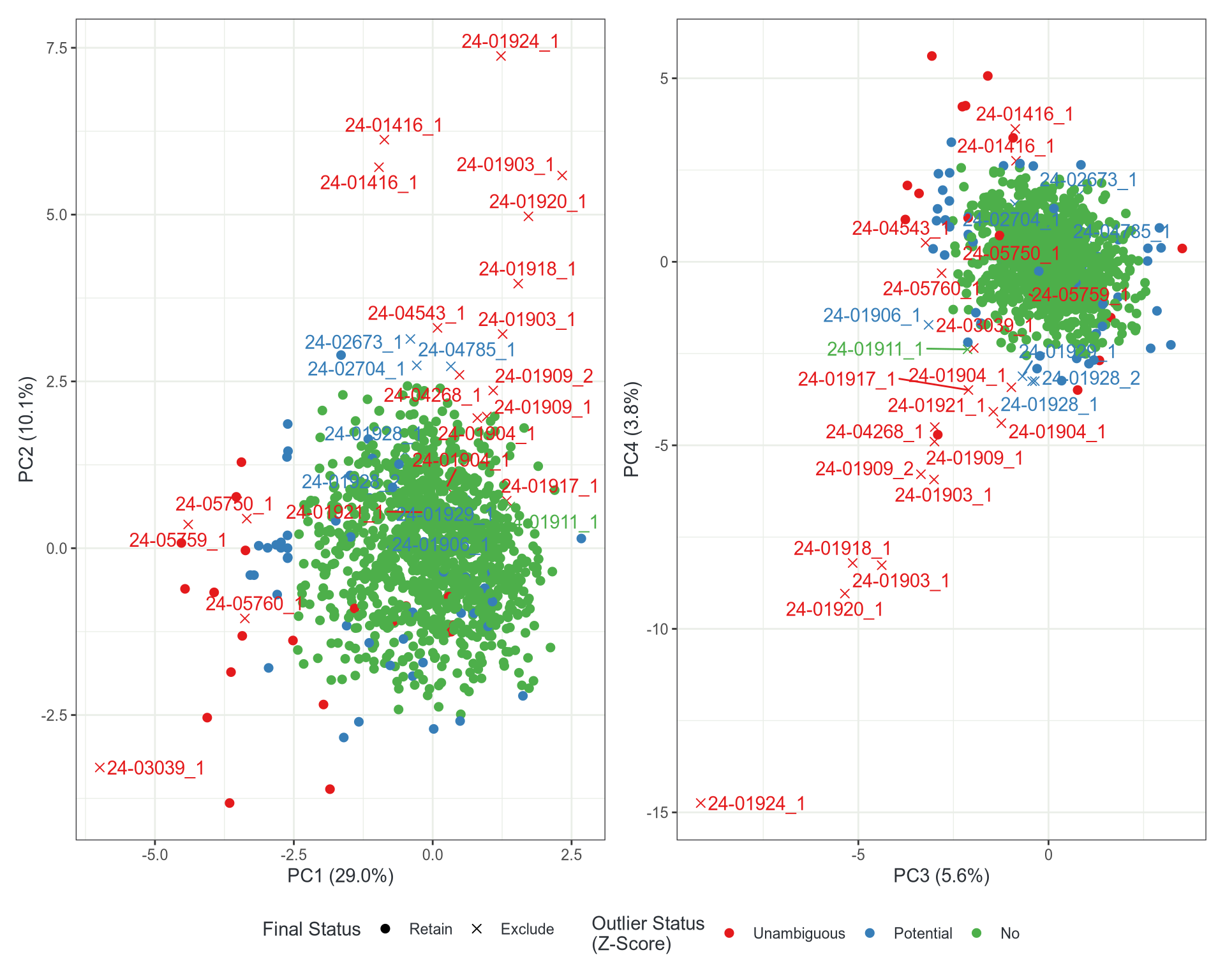
Normalisation
- Do we have batch effects?
hRUV(Kim et al. 2021),RUV(Risso et al. 2014),RUVIII-NB(Salim et al. 2022)- TMM (Robinson and Oshlack 2010),
CQN(Hansen, Irizarry, and Wu 2012)
hRUVhard-wired to USyd metabolomics structure- Rewrote significant amount of (github-only) package
RUVIII-NBalso needed rewriting (github only)
- Didn’t need them anyway 😅
Batch Effects
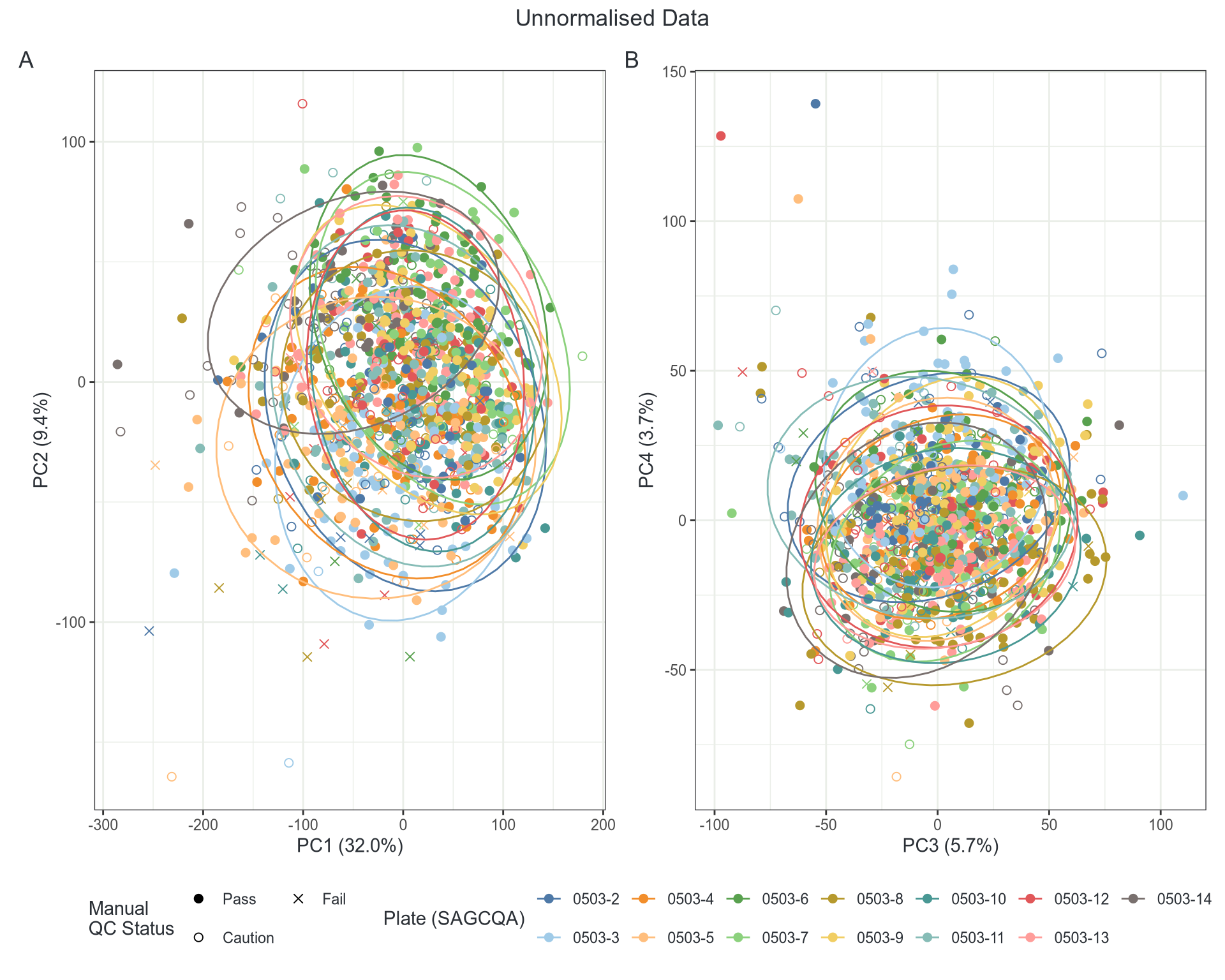
GC & Length Artefacts
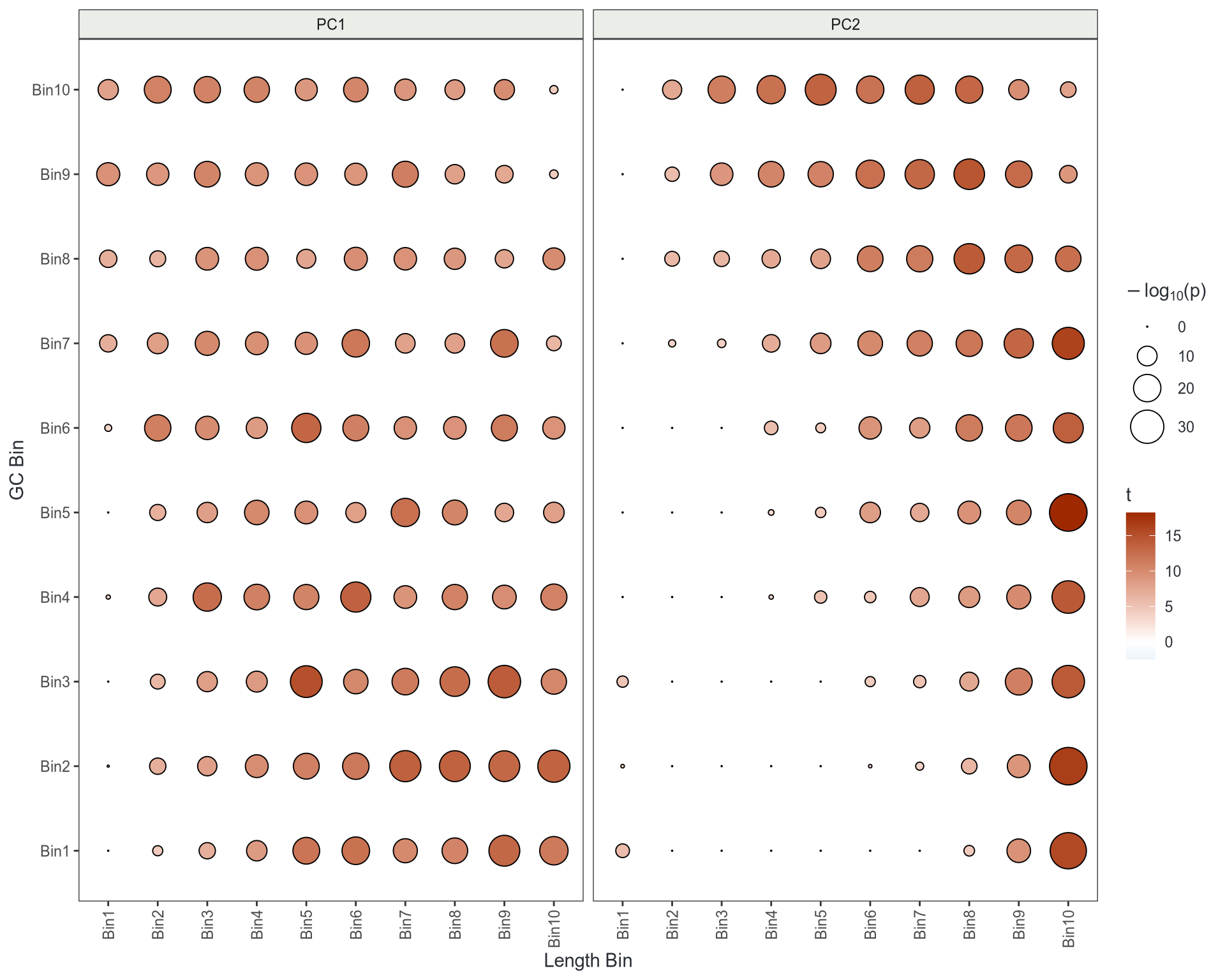
- Method developed by Lachlan Baer
- Bin gene-level rotations from PCA by GC & Length
- Perform \(t\)-test in each bin
- If no bias \(\implies\) random noise
- Looks like GC & length is impacting signal
Conditional Quantile Normalisation
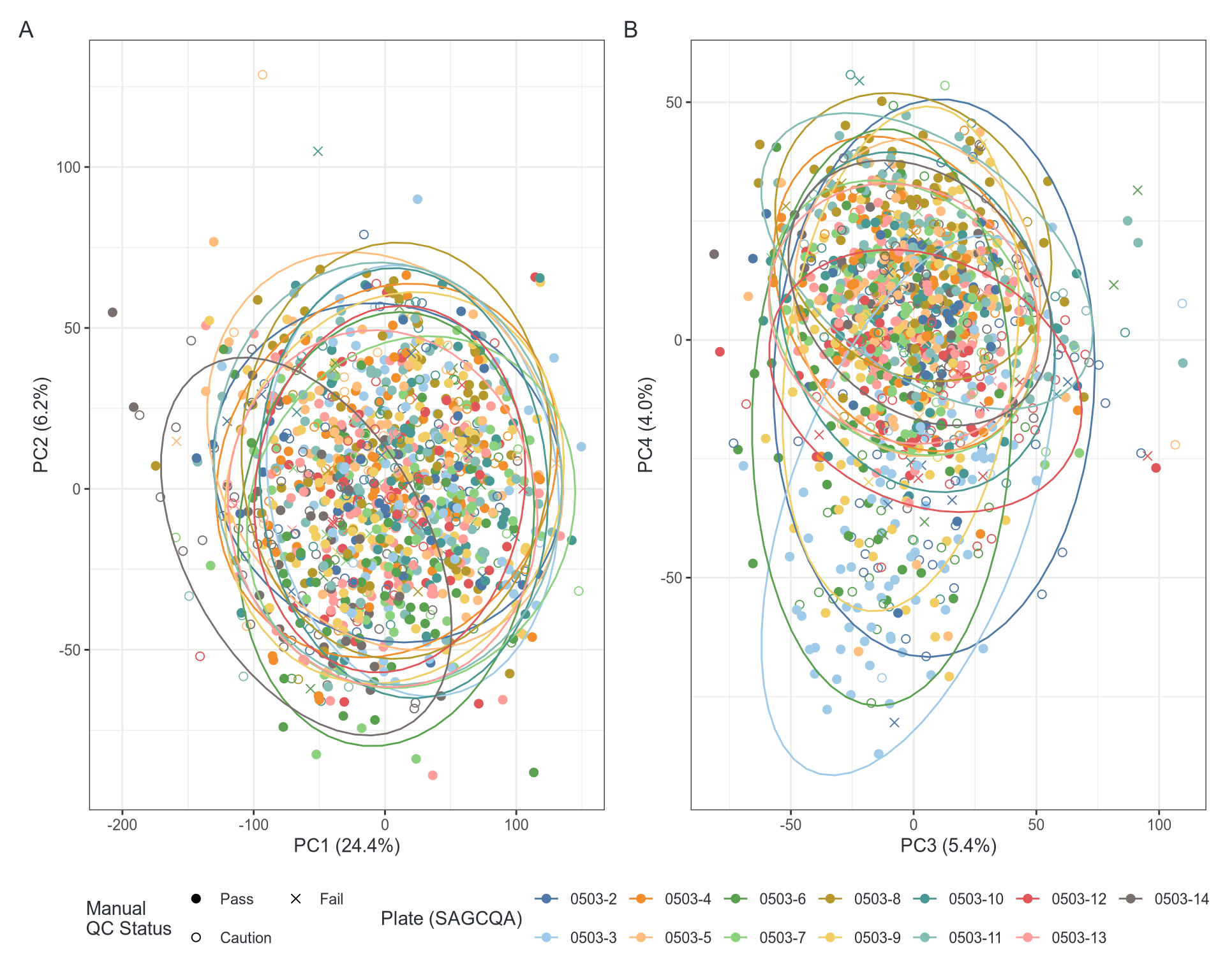
PC4 captures age 🎉
Performance on Pools
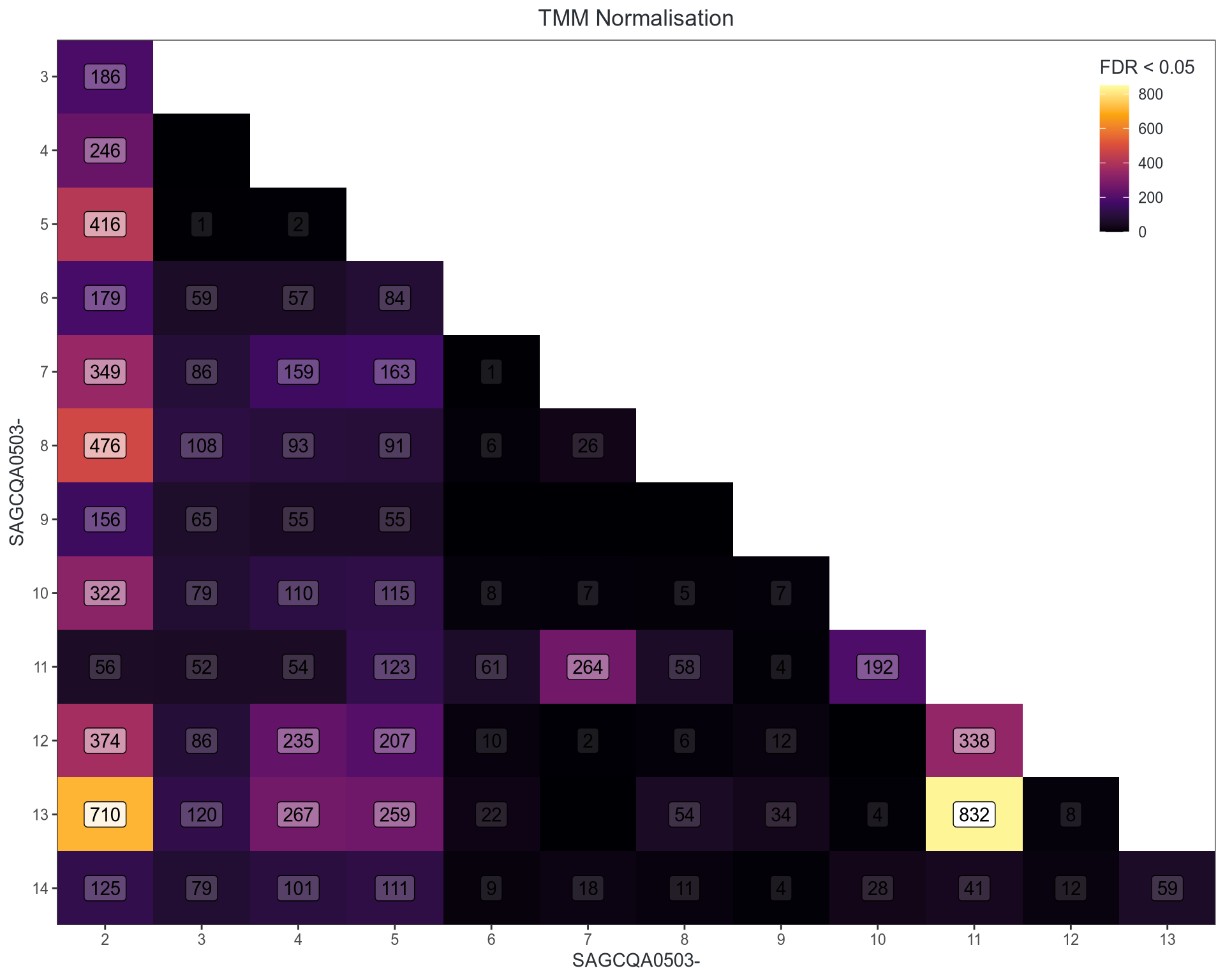
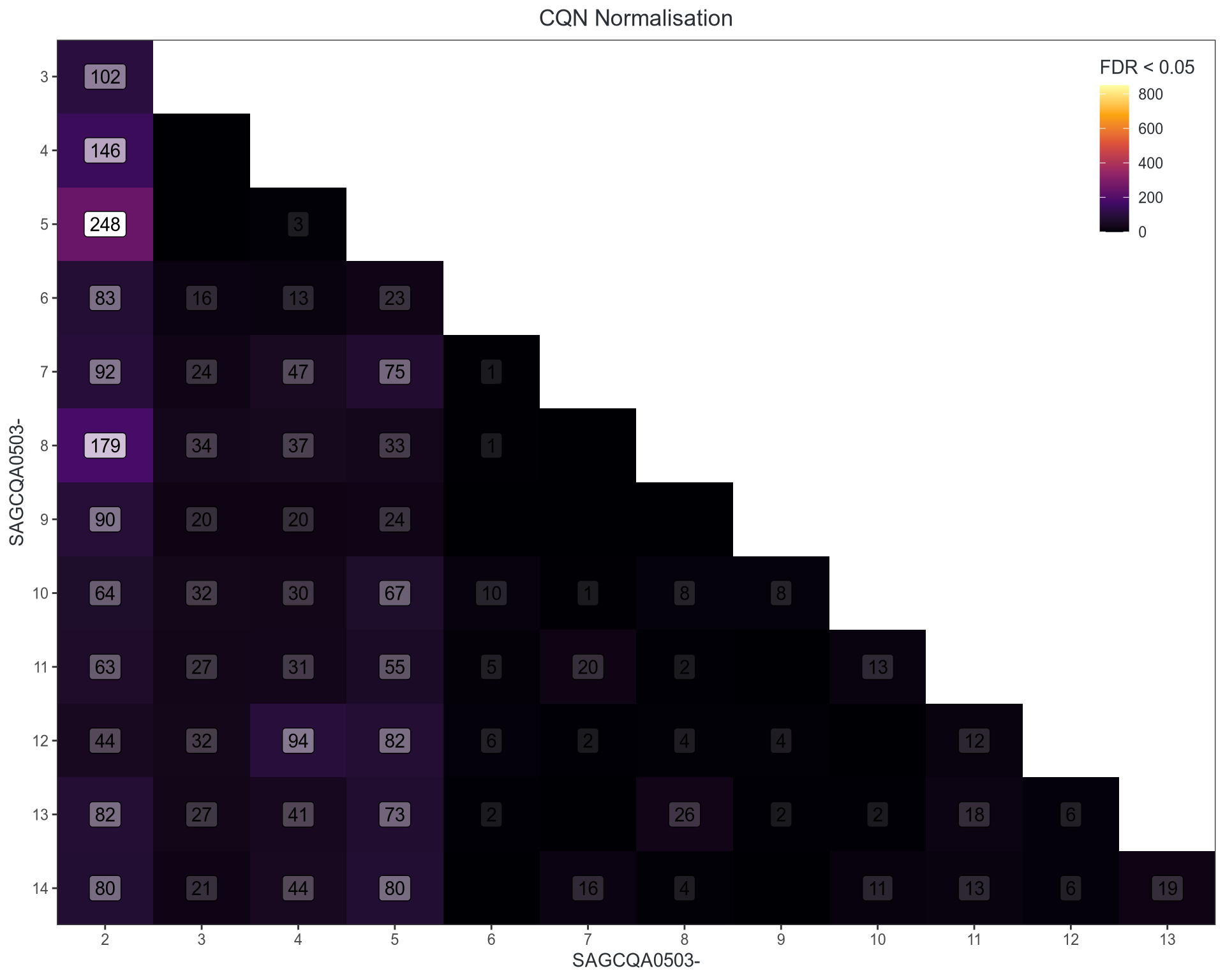
Both improve further using glmTreat() at FC > 1.1
Do Any QC Metrics Impact Signal
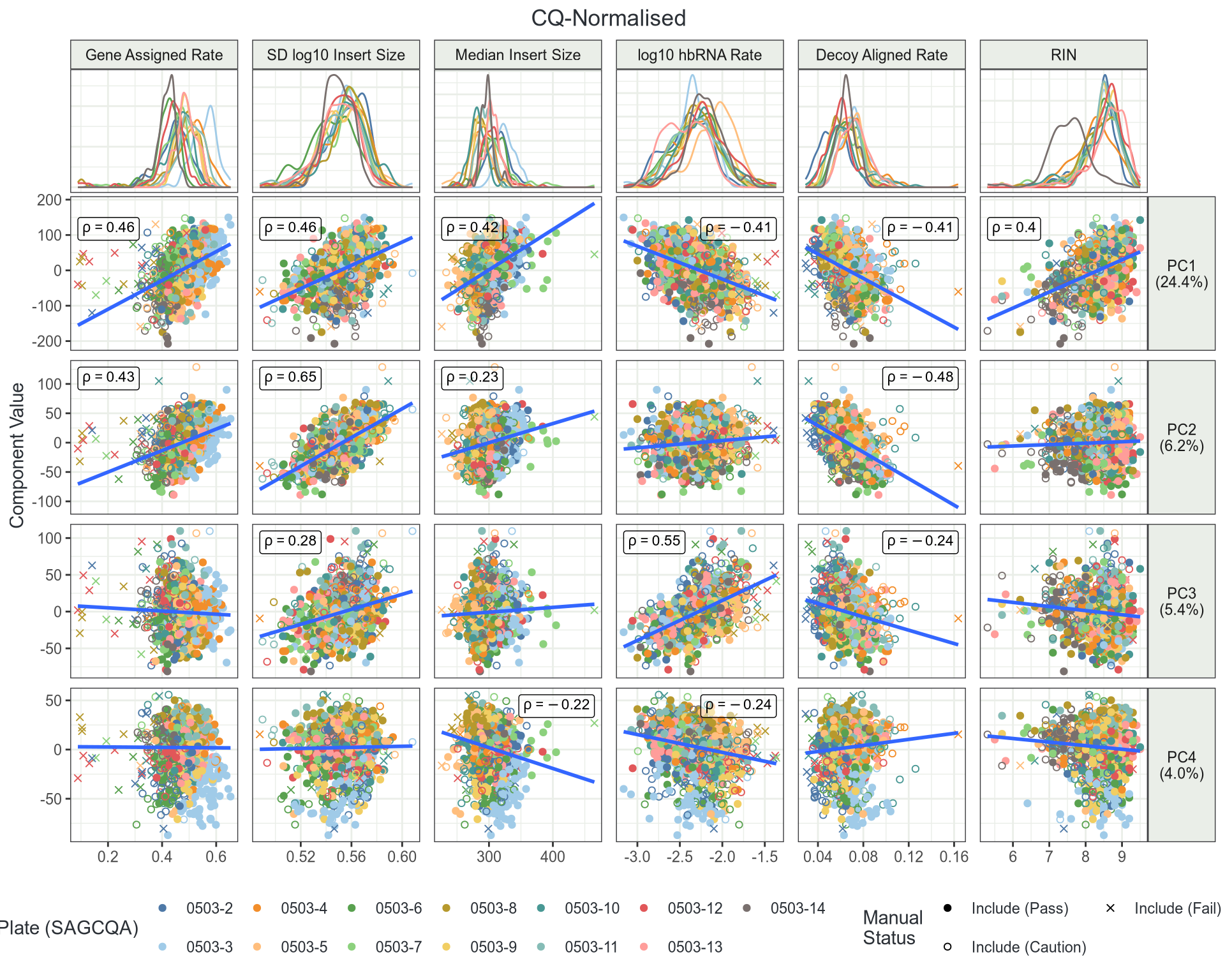
- Several QC metrics strongly correlate with PC1-3
- How will this impact DGE analysis?
- Can I check?
Using QC Metrics As Predictors
Managing Noise
- Strategy is to fit a null (intercept-only) model
- Add QC metrics to find which improve fit
\(\implies\) becomes baseline model for all DGE analysis - Identify best combination of metrics
- Add QC metrics to find which improve fit
- gDNA-related metrics (QC PC1)
stranded_coverage,prop_no_feature,sd_log10_insert_size,decoy_aligned_rate,gene_assigned_rate
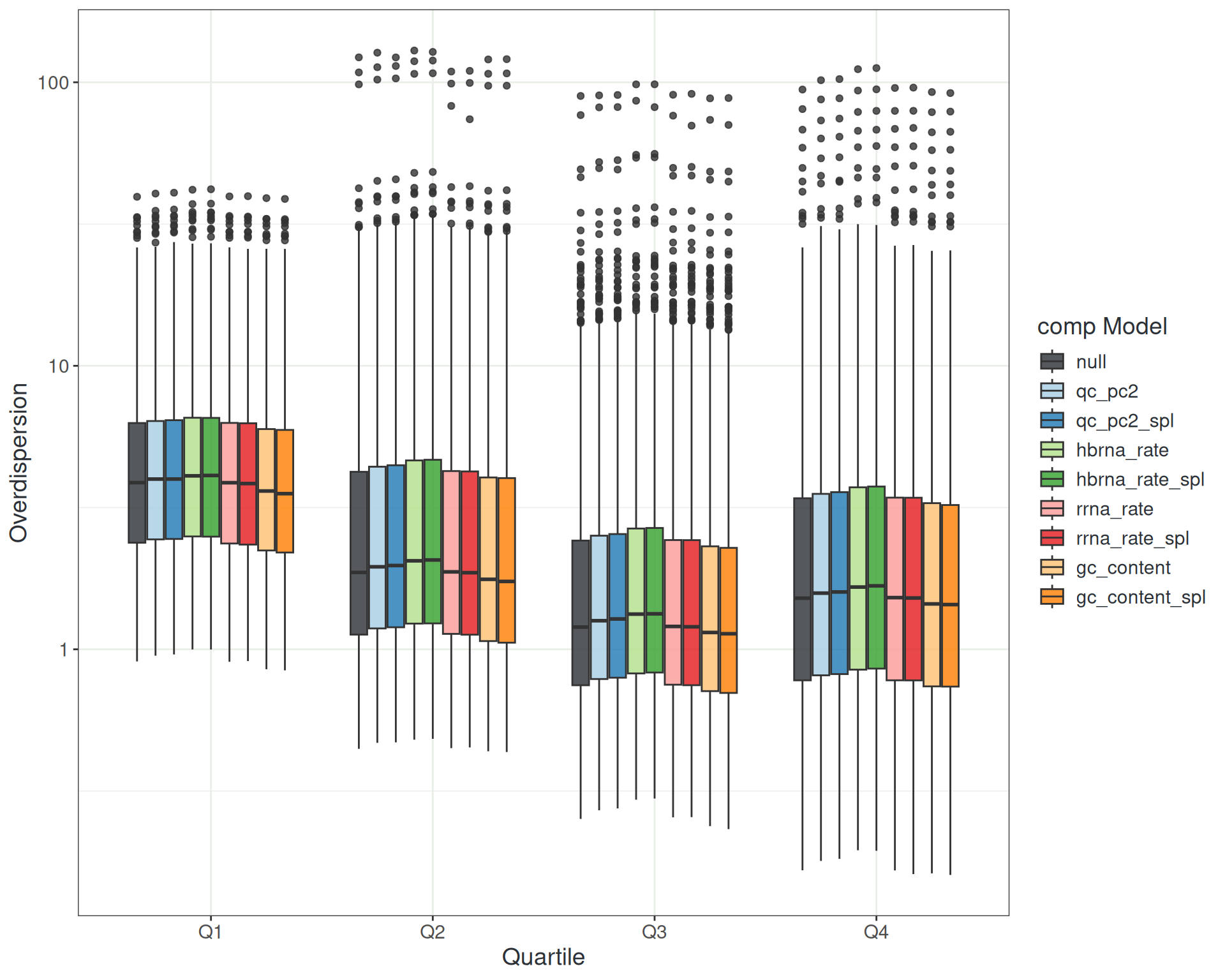
- Library composition metrics (QC PC2)
hbrna_rate,rrna_rate,gc_content,duplication_rate
- Also tried RIN
Managing Noise: gDNA Metrics
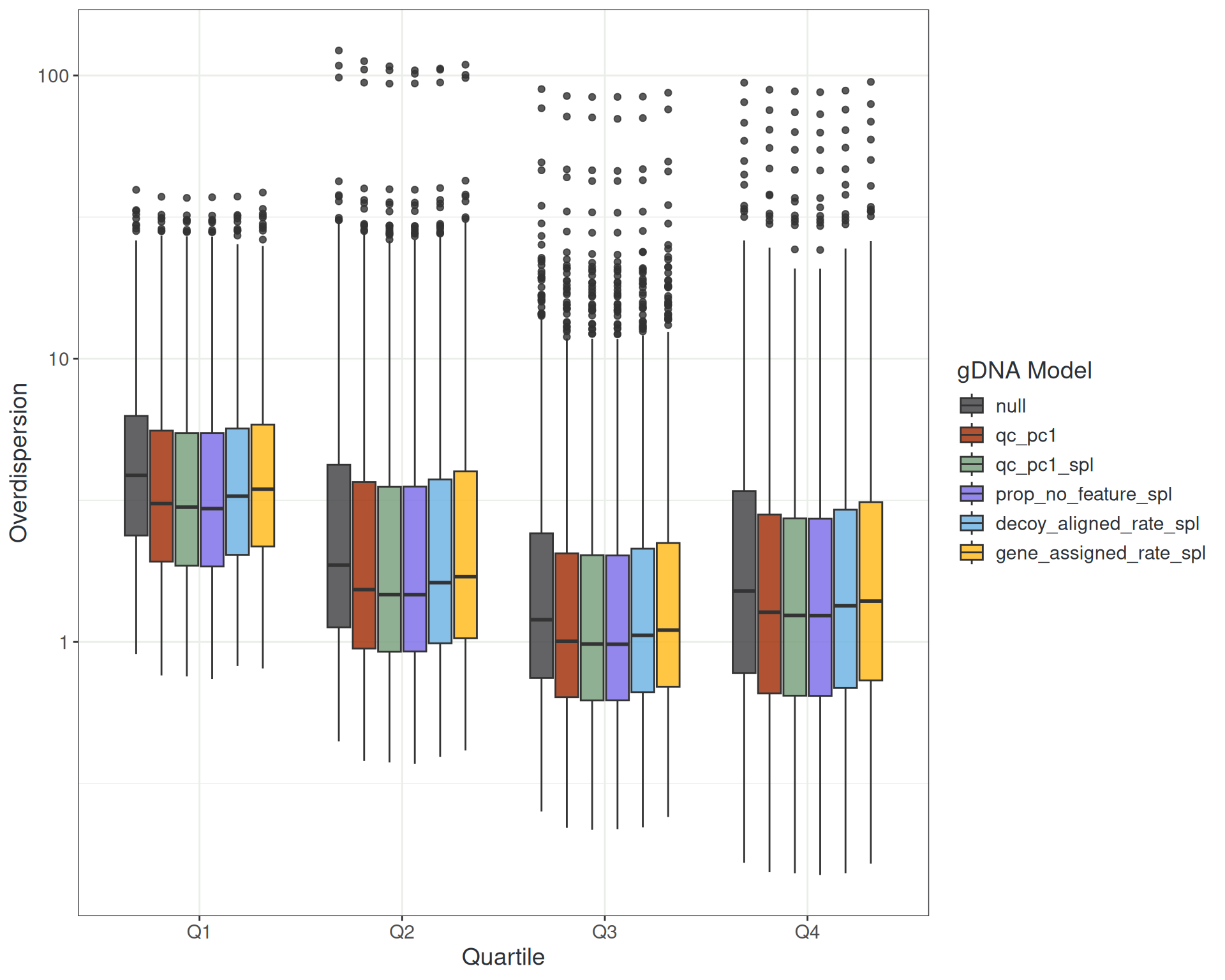
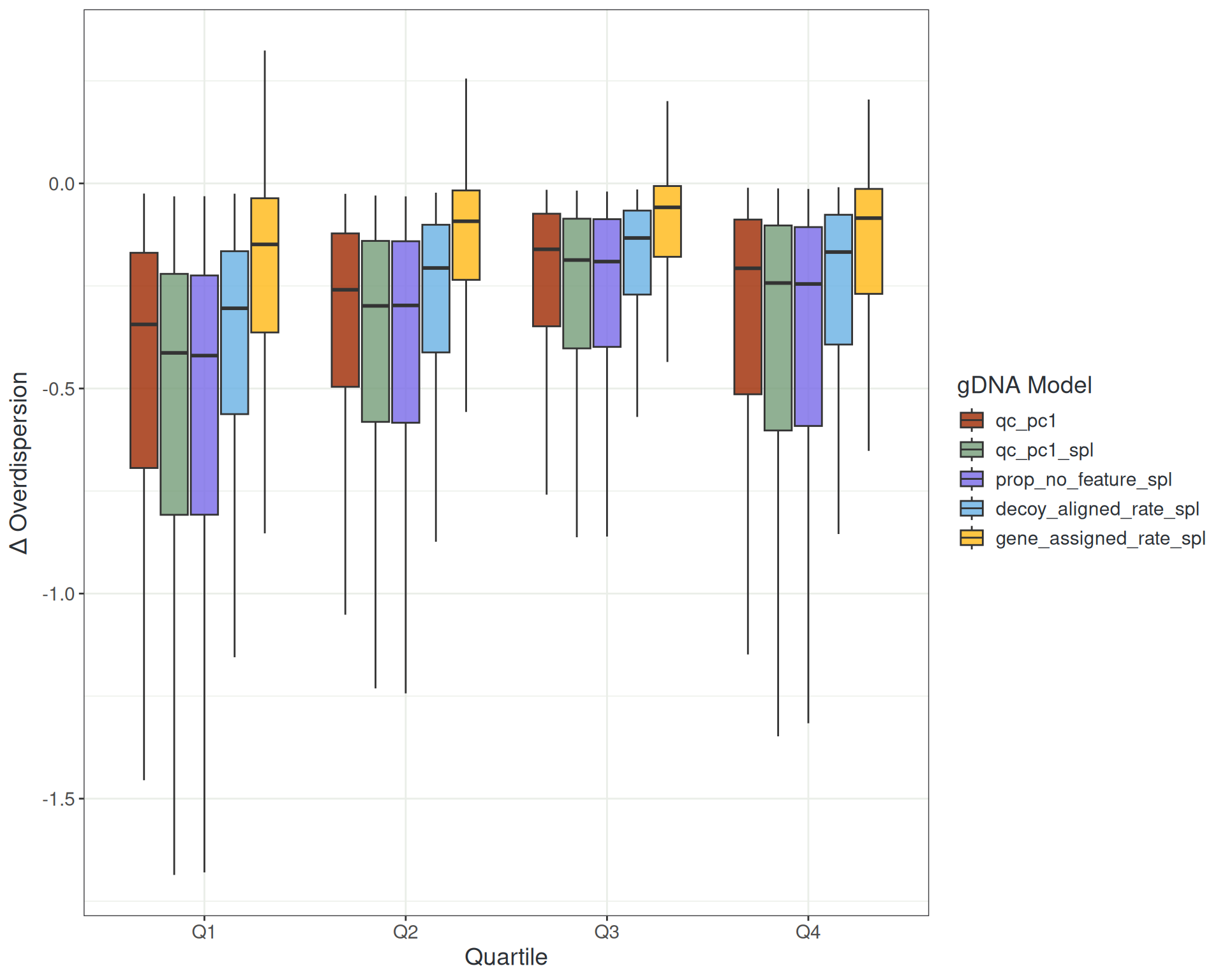
Managing Noise: Library Composition
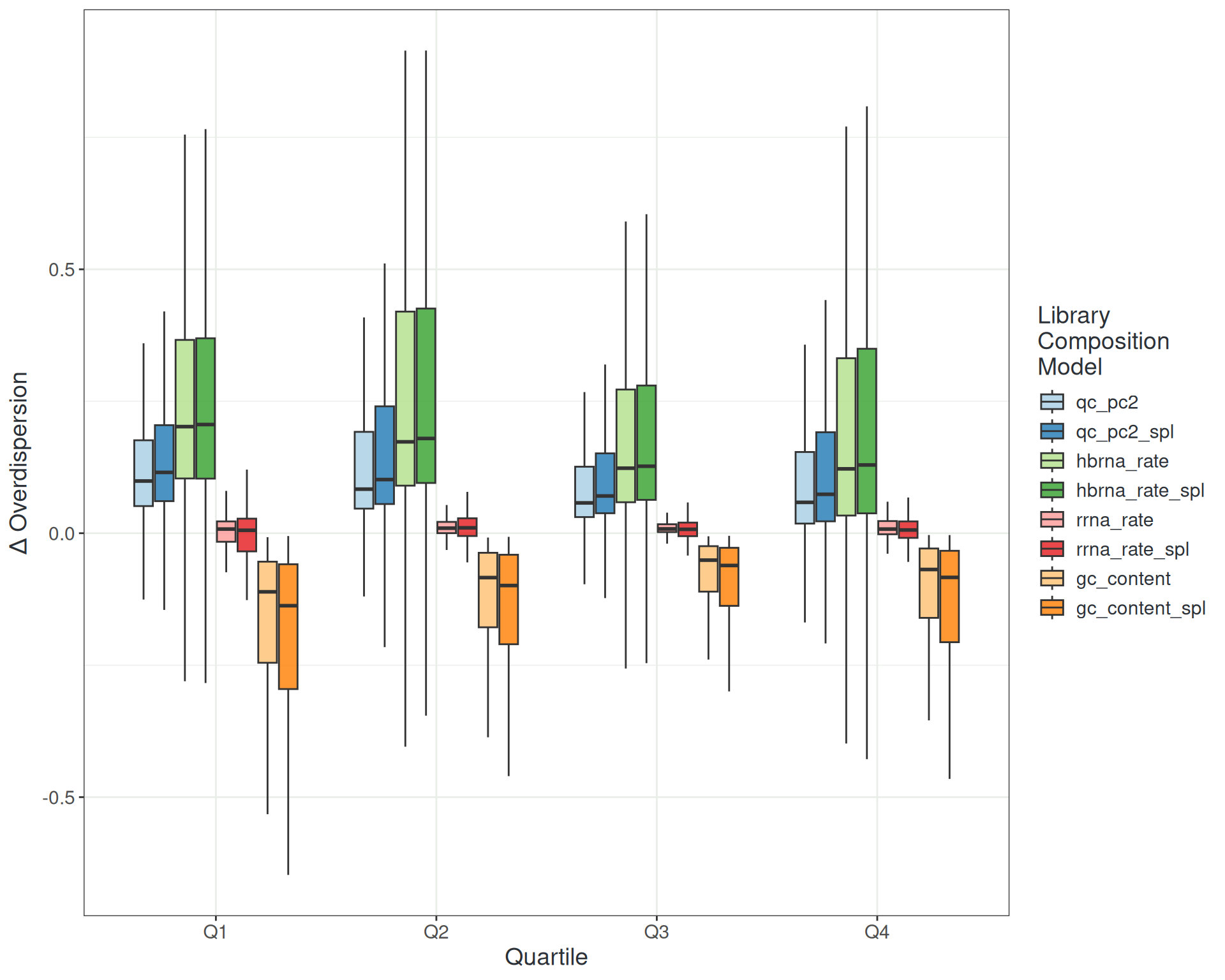
Managing Noise: Combined Metrics
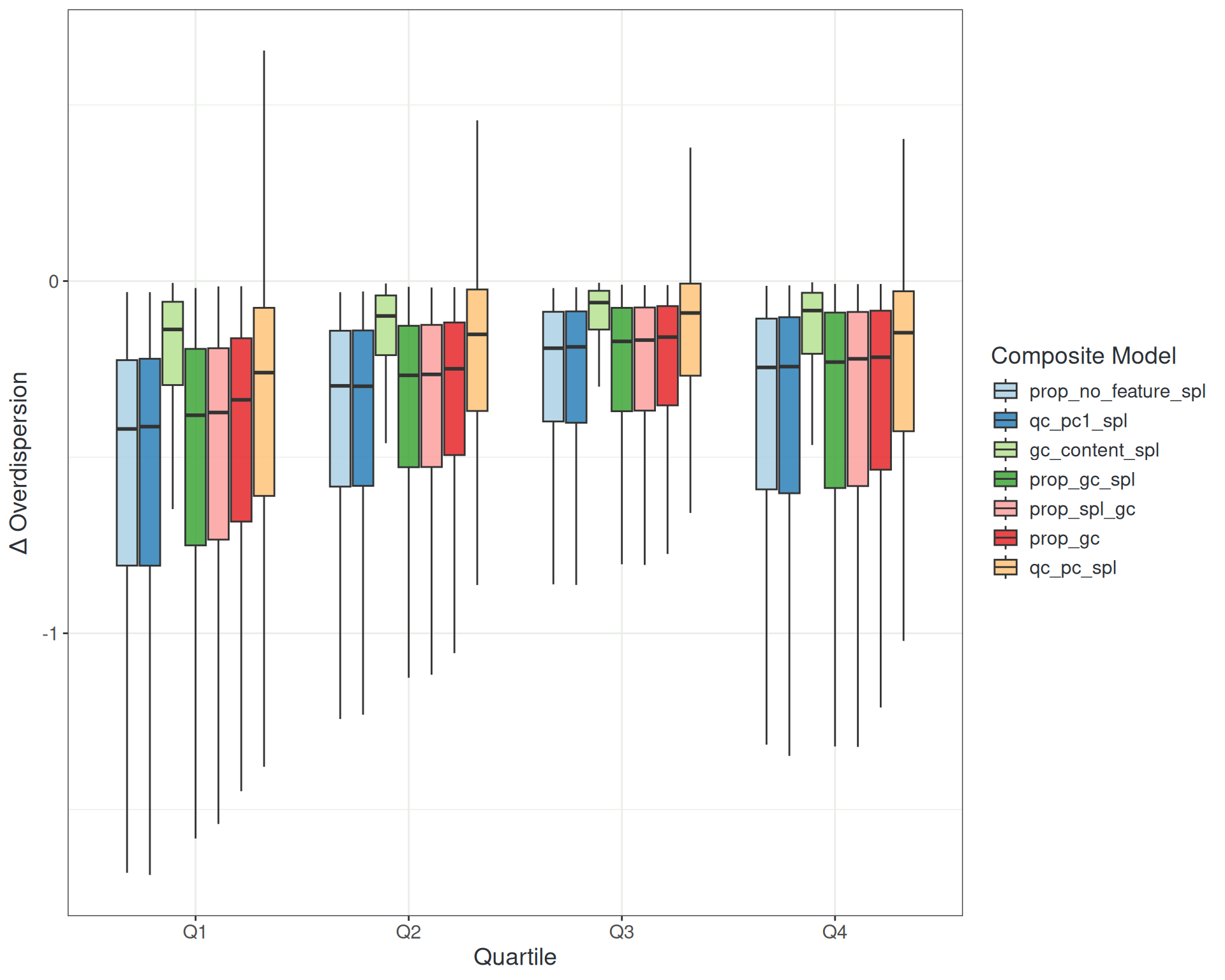
- No combination outperformed
prop_no_featureas a natural spline- Each predictor costs degrees of freedom
- Have also tested age as a predictor
- Natural spline is better than linear
- PC1 from genotype data improves about 30% of genes
Take Home
- Unlike any dataset I’ve ever worked with
- The gDNA aspect reinforces Hien To’s work on
strandcheckR- Would love to see it added to
nf-core
- Would love to see it added to
- Setting all data handling early is great
- Unexpected challenges incorporating mixups
- Typos in sample sheets can be infuriating (e.g. Pool vs pool)
- Didn’t see any clinical variables for >2yrs
- T2D: 000’s of DE Genes
- CKD: 000’s of DE Genes
- CHD is a really difficult phenotype
Clickbait Time
Is large cohort transcriptomics just scRNA without the zero counts?

The questions are completely different:
- We want clusters of genes defining disease pathology with age + complication, NOT clusters of cells
- QC represents fundamentally different technical processes
Acknowledgements
Black Ochre Data Labs
Alex Brown
Jimmy Breen
Alastair Ludington
Sam Buckberry
Liza Kretzschmar
Yassine Souilmi
Katharine Brown
Rose Senesi
Sam Godwin
Rebecca Simpson
Adam Heterick
Kaashifah Bruce
Bastien Llamas
Amanda Richards-Satour
Justine Clark
Holly Massacci
Sarah Munns
Alli Foster
Sehaj Dhariwal
SAGC
Sen Wang
Renee Smith
John Salamon
Paul Wang
Baker Heart & Diabetes Institute
Sam El Osta
Ishant Kurma
Moshe Olshansky
Scott Maxwell
SAHMRI / Wardliparingga
Natasha Howard
Victor Chang Cardiac Research Institute
Jason Kovacic
University of Sydney
Jean Yang
Centre For Population Genomics
Dan MacArthur
References
