Alternate References For Transcriptomics
Adelaide Bioinformatics Seminars
Black Ochre Data Labs / ANU
May 3, 2024
Acknowledgement of Country
I would like to acknowledge that many of us are meeting today on Kaurna Country.
I acknowledge the deep feelings of attachment and relationship of the Kaurna people to their Place.
I also pay my respects to the cultural authority of Aboriginal and Torres Strait Islander peoples from other areas of Australia online today, and pay my respects to Elders past, present and emerging.
About Me
- Postdoctoral Fellow, Black Ochre Data Labs, Indigenous Genomics, Telethon Kids Institute
- 2020-2022: Dame Roma Mitchell Cancer Research Labs, University of Adelaide
- 2014-2020: Bioinformatics Hub, University of Adelaide
- 2008-2018: Slowest PhD Student in the world…
- Dec 2002: First used R &
limma
- 1st Year B.Sc. in 1986 \(\implies\) Dropped out in 1987




Bioconductor Package Developer



![]()

motifTestR
- Was adding motif enrichment testing to the GRAVI workflow
- Decided to avoid the MEME-Suite 🤯
- Plays badly with
conda - I struggle to interpret the results
- Not R native (although
memesdoes wrap some of it)
- Plays badly with
![]()
- Particularly interested in motif positions within a set of sequence
- Analogous to
centrimo
- Analogous to
- Added enrichment testing because I could
motifTestR: Positional Bias
- If no positional bias \(\implies\) motifs will match uniformly across the sequence width
- Break sequences into bins
- Assign probabilities (\(\pi_i\)) of matches within bins \(i = 1, ... I\)
- Proportion of the total viable positions for a match within each bin
- Test matches in bin \(i\) are as expected using \(\pi_i\) \(\implies p_i\)
binomial.test()
- Use harmonic mean p-value to obtain a single p-value for each motif
- Returns ‘enriched sequence regions’ where \(p_i\) < \(HMP(p.)\)
motifTestR: Positional Bias
- Can be performed using:
- absolute distance from centre or
- the entire sequence width
- All sequences must be the same length
- Relies on a single ‘best match’ per sequence (but also applies weights)
- Also finds motif matches for you
- Heavy reliance on
Biostrings
- 7,000 seq \(\times\) 400bp \(\times\) 400 motifs \(\implies\) < 30sec (phoenix: 8 cores)
- 50,000 seq \(\times\) 800bp \(\times\) 400 motifs \(\implies\) ~2min (phoenix: 8 cores)
motifTestR: Positional Bias Plots
A <- top_matches %>%
plotMatchPos(
se = FALSE, abs = TRUE, linewidth = 1/2,
binwidth = 5
) +
labs(
x = "Distance From Centre",
y = "Proportion",
colour = "Motifs In Cluster"
) +
theme(legend.position = "none")
B <- top_matches %>%
plotMatchPos(
type = "cdf", geom = "line", abs = TRUE,
linewidth = 1 / 2, binwidth = 1
) +
labs(
x = "Distance From Centre",
y = "Proportion",
colour = "Representative Motif"
)
A + B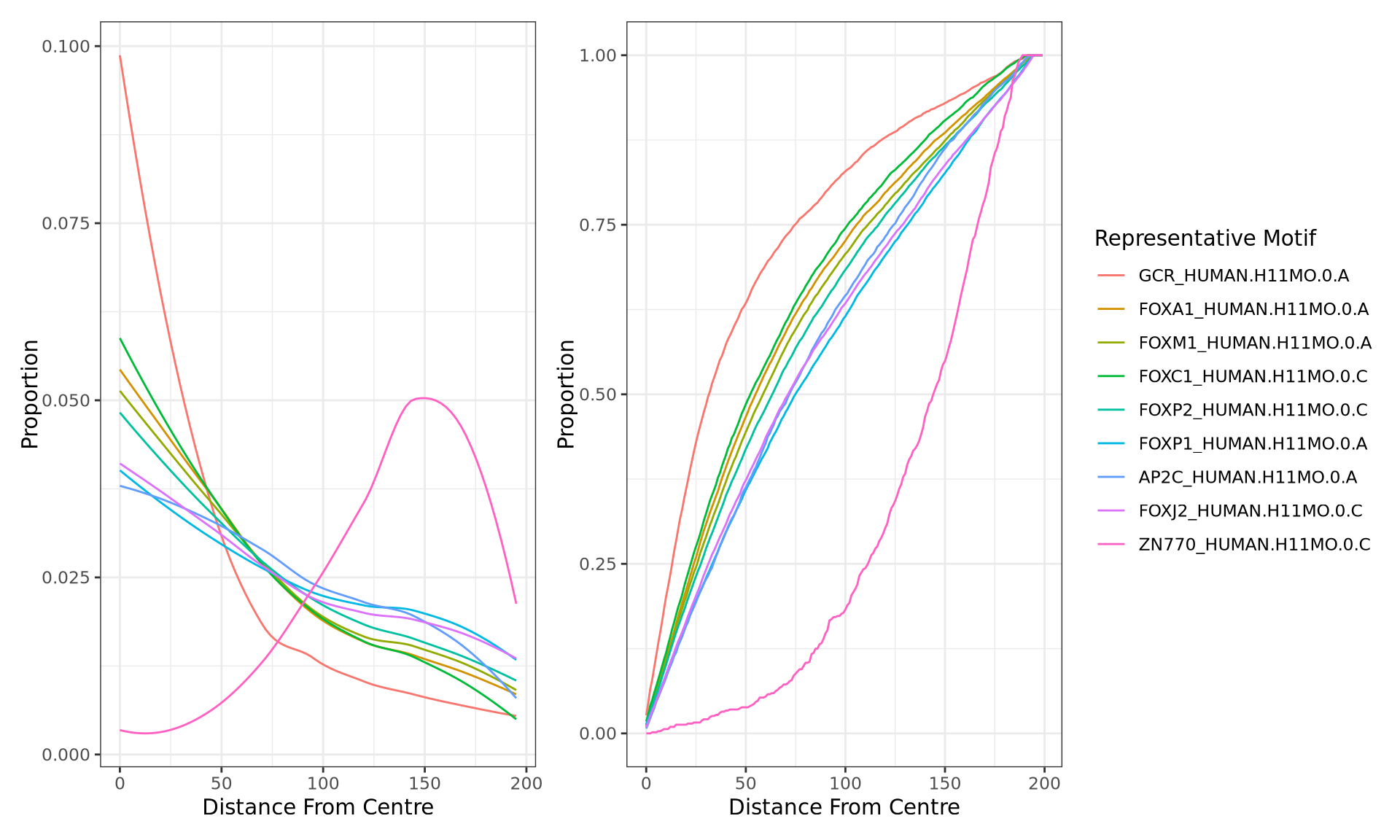
motifTestR: Positional Bias Plots
top_matches %>%
plotMatchPos(
geom = "point", abs =TRUE, use_totals = TRUE,
binwidth = 1
) +
geom_smooth(
se = FALSE, colour = "black",
linewidth = 1/2, method = 'loess'
) +
facet_wrap(~name, labeller = as_labeller(lb)) +
theme(legend.position = "none") +
labs(
x = "Distance From Sequence Centre",
y = "Total Matches"
)Show the matches at each position
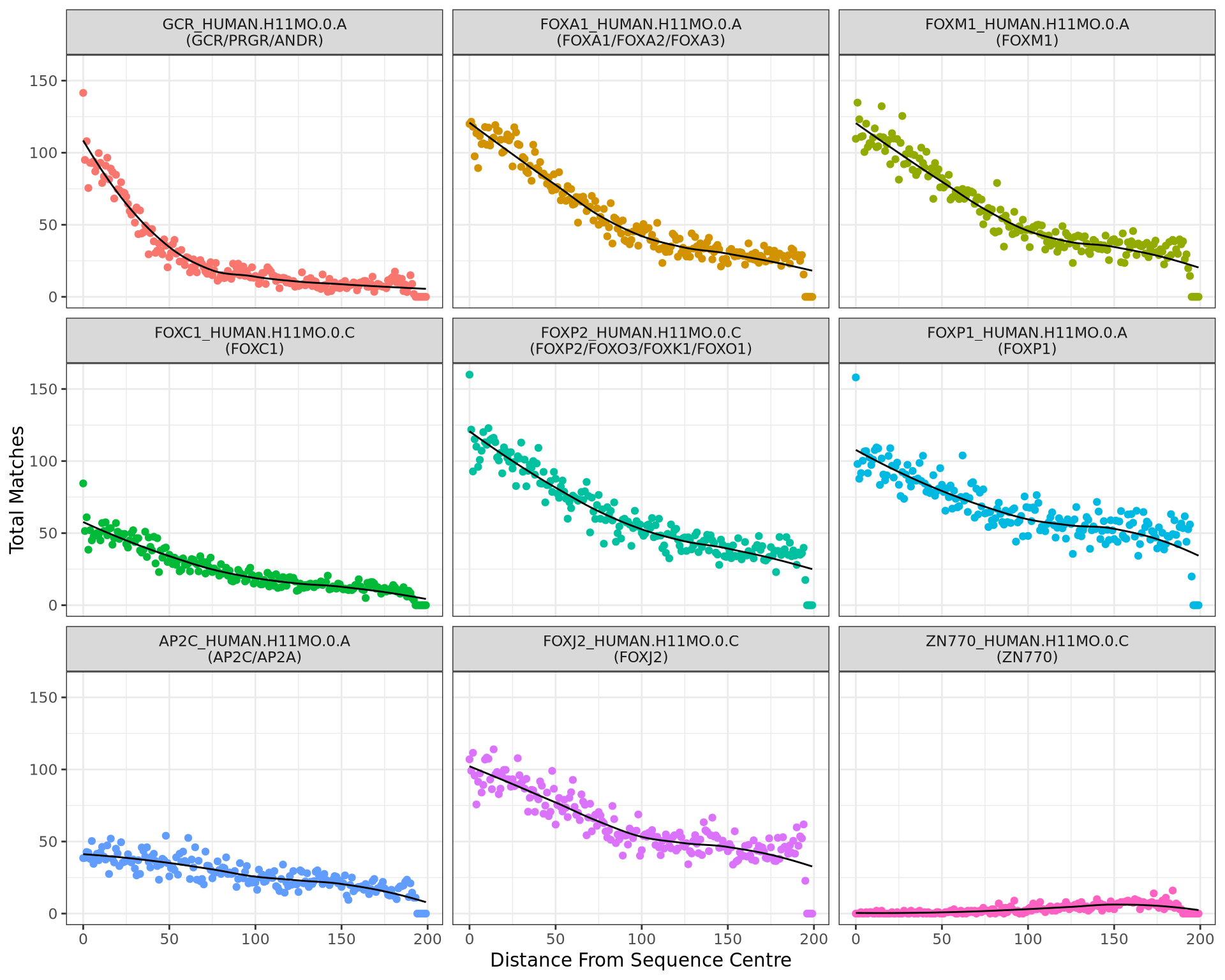
motifTestR: Positional Bias Plots
motif_list %>%
dplyr::filter(altname %in% top_pos) %>%
to_list() %>%
getPwmMatches(
fw_seq, abs = TRUE, best_only = TRUE,
mc.cores = threads
) %>%
plotMatchPos(
abs = TRUE, type = "heatmap",
cluster = TRUE, use_totals = TRUE,
binwidth = 10
) +
labs(
x = "Bin", fill = "Total\n Matches"
)- Clustering & heatmaps can help identify redundancy
- Useful for larger sets of motifs / matches
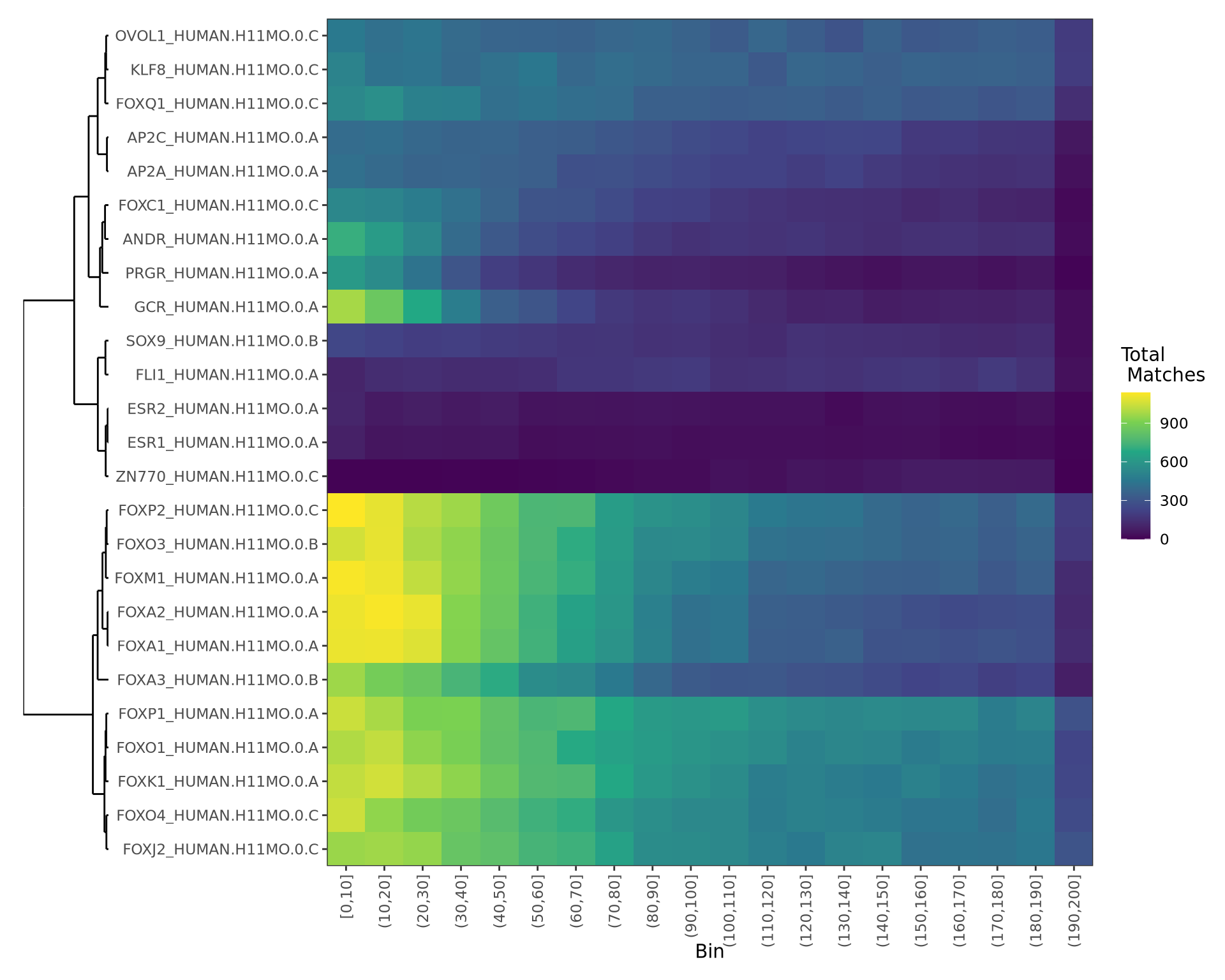
motifTestR: Motif Enrichment
- Hypergeometric: Subset A vs Subset B
- Specifically for sequences where one subset changes & the other doesn’t
- Realistically, results are specific to the each BG set
- Poisson: SetA Vs BG
- BG Set >>> Set A
- Estimates match rate per sequence in BG \(\implies\)
poisson.test() - Quick(ish) & dirty
- Explicitly assumes matches are Poisson
- QuasiPoisson: SetA Vs BG (In Blocks)
- Tests match rate per block (i.e. iteration) in BG
- Allows for overdispersion
- Iterations: SetA Vs BG (In Blocks)
- No distributional assumptions (except CLT)
- Derives mean & sd per iteration in BG
\(\implies\) \(Z\)-score + p-value
motifTestR: makeRMRanges()
- Added function for obtaining Random Matched Ranges for a test set
- Relies on a discrete set of features
- e.g. promoters, enhancers, etc
- Samples RMRanges to exactly match the feature distribution of the test set
- Can be sampled in blocks for iterative methods (quasipoisson / iteration)
motifTestR: Motif Enrichment
- Is much, much slower… 😞
- 7000 \(\times\) 400bp \(\times\) 400 motifs (quasipoisson):
- Using 7000 \(\times\) 1000 = 7x106 RMRanges
- 7 minutes (phoenix: 8 cores)
- 50,000 \(\times\) 800bp \(\times\) 400 motifs (quasipoisson):
- Using 50,000 \(\times\) 1000 = 5x107 RMRanges
- 1hr 35min minutes (phoenix: 8 cores)
- 400 Vs 6500 \(\times\) 400bp \(\times\) 400 motifs (hypergeometric):
- 35 sec (phoenix: 8 cores)
And Now My Real Talk…
Current Research
- PROPHECY (Preventing Renal OPthalmic and Heart Events in CommunitY)
- Multi-Omics study led by Prof Alex Brown, with Chief Data Scientist A/Prof Jimmy Breen
- Investigating the very high rates of Diabetes within Indigenous Australians
- Also very high rates of complications: CVD & CKD
- Is strongly led by Community in partnership with BODL
- Hoping to bring benefits of Precision Medicine
- New approach to Indigenous Health \(\implies\) listen first
Life Expectancy Gaps
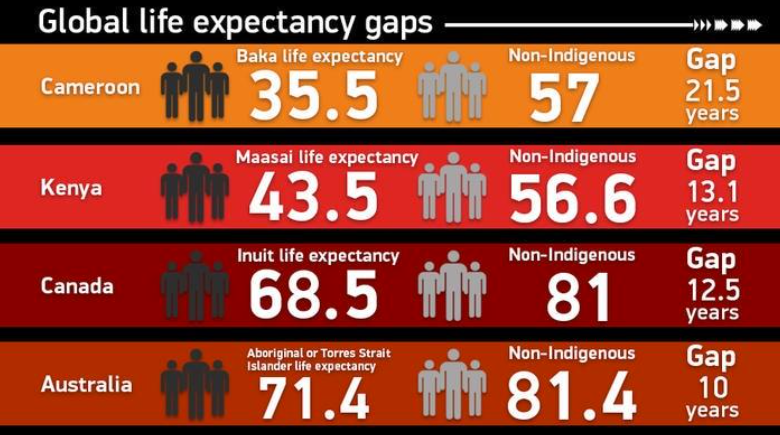
Courtesy of A/Prof Jimmy Breen, Black Ochre Data Labs, TKI
Mortality From Diabetes
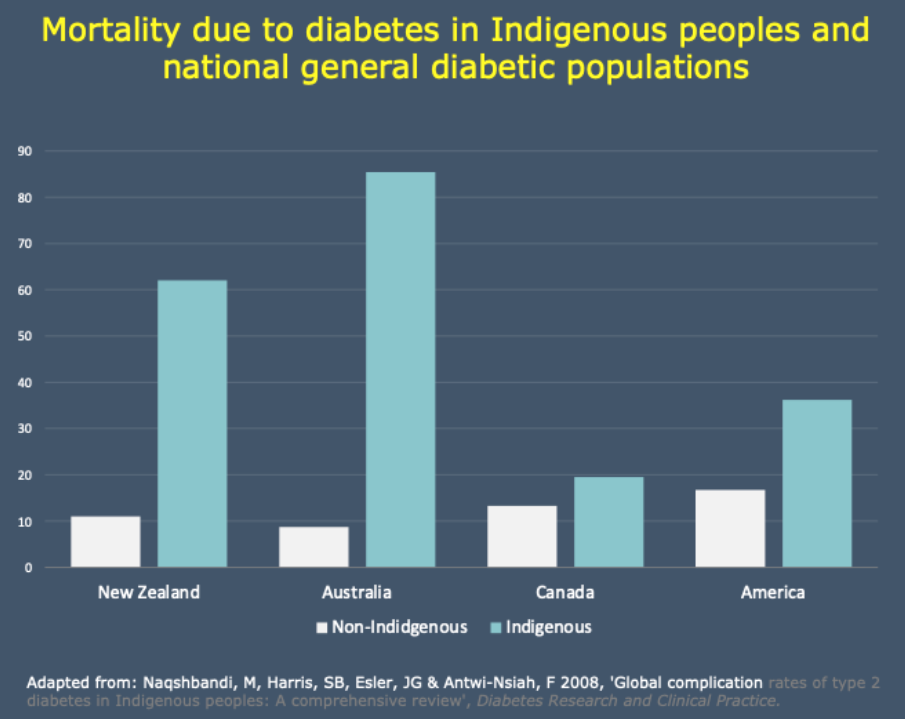
Why?
- Taking all socio-economic indicators \(\implies\) disease risk is still hugely increased for Indigenous Australians
- Not only disease risk, but higher rates of profound complications
- Are there risk alleles?
- If so, what are they? How do they contribute to risk?
- Are there protective alleles?
- Is there an epigenetic contribution?
- Additional environmental factors?
Multi-Omic Approaches
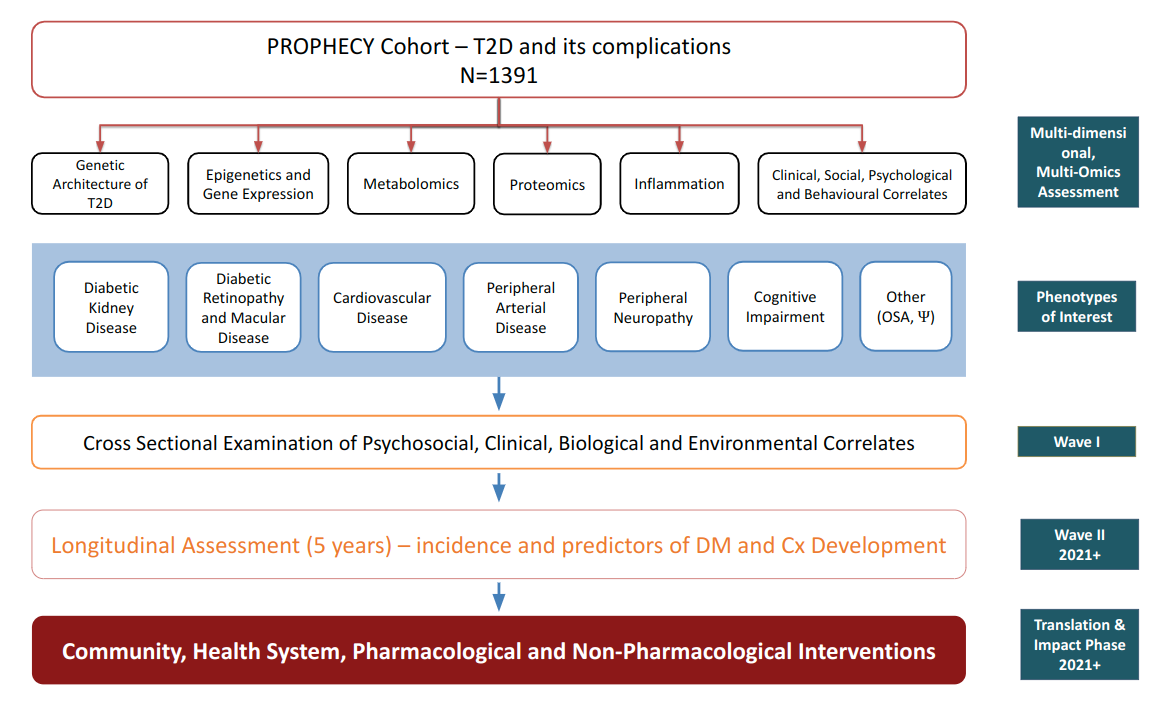
The Transcriptomics Layer
- Whole blood samples
- Can we capture dynamics of disease / signatures / biomarkers etc?
- Approach will be classical statistics + network approaches
- Tried to be excruciatingly careful with design:
- plate layout, pooled replicates, globin-depletion, ERCC spike-ins, RIN scores etc
- Library preps are finally underway
- Data lands around September
Why Think About Reference Genomes?
Genetic Diversity
- Indigenous Australians are almost totally absent from repositories of genetic variation
- \(\sim\) 25% of variation within Indigenous Australian populations is unique & poorly characterised (Easteal et al. 2020)
- If contributing to disease risk and we use a standard reference \(\implies\) will we see it?
- Can we bring the reference closer to our participants’ genomes?
- NCIG are very close to a pangenome graph assembled from Indigenous Australians
- A little too far off of to be viable yet
- Monica Guilhaus doing an amazing job figuring out spliced pan-transcriptome graphs (Sibbesen et al. 2023)
Gene-Level Approaches
- STARconsensus (Kaminow et al. 2022) enables inclusion of a population-specific consensus variant set
- Also possible for personalised genomes
- Now has a
diploidmode for phased, personalised variant sets
- Incorporates variants during indexing
- Indexes twice (Standard + Modified)
- Returns alignment co-ordinates relative to the unmodified reference
- Simple to obtain counts using standard methods (e.g. featureCounts)
The Downside
- Locked into using STAR
- In my hands can’t align full-length, reference-derived transcripts
- Counts are gene-level
- We understand the distribution of counts at the gene level
edgeR&DESeq2: well established Negative Binomial methods
- Transcript-level counts have additional uncertainty
- Need to be scaled by an over-dispersion estimate
\(\implies\) can then be treated as Negative Binomial (Baldoni et al. 2024) - Cannot obtain these from
STAR+featureCounts
- Need to be scaled by an over-dispersion estimate
Transcript-Level Analysis
salmonprovides transcript-level counts with overdispersion estimates!- Can we modify a reference transcriptome using a set of variants?
transmogR + salmon
- Takes about 15-20 mins, even on a laptop
- Modified transcripts can be optionally tagged:
- Indicating variant types (
s/i/d) - Naming the variant set (e.g.
panhuman)
- Indicating variant types (
- Generally using SNVs + InDels < 50bp
- I’ve been using a modified transcriptome & modified genome as the gentrome
- The genome is used by
salmonas decoy sequences (Srivastava et al. 2020)
- The genome is used by
- Indexing for
salmonstill takes a while…
DBPAInT: Developing Best Practices for the Analysis of Indigenous Transcriptomes
The Approach
- STARconsensus: using variants from the wrong population still improves performance (Kaminow et al. 2022)
- 1000 Genomes Project Panhuman
- alleles found in >50% of unrelated across entire project
- 1000 GP AFR Subpopulation
- alleles found in >50% of unrelated within the AFR population
- most distinct population from the Panhuman
- PROPHECY consensus set
- alleles found in > 50% of unrelated members of the study
- Used 3 familial steps to define ‘unrelated’
The Analysis
- 6 female pilot samples
- All genomes are y-masked
- Replicating elements of STARconsensus
- Taking GRCh38 as the worst case \(\implies\) changes are improvements?
- Four sets of genomic alignments / sample
- Personalised variants arriving in the next few days (weeks?)
- Also running a
transmogR+salmonanalysis- Four sets of transcriptomic alignments / sample
- Haven’t figured out how to do a diploid transcriptome yet…
The Variants
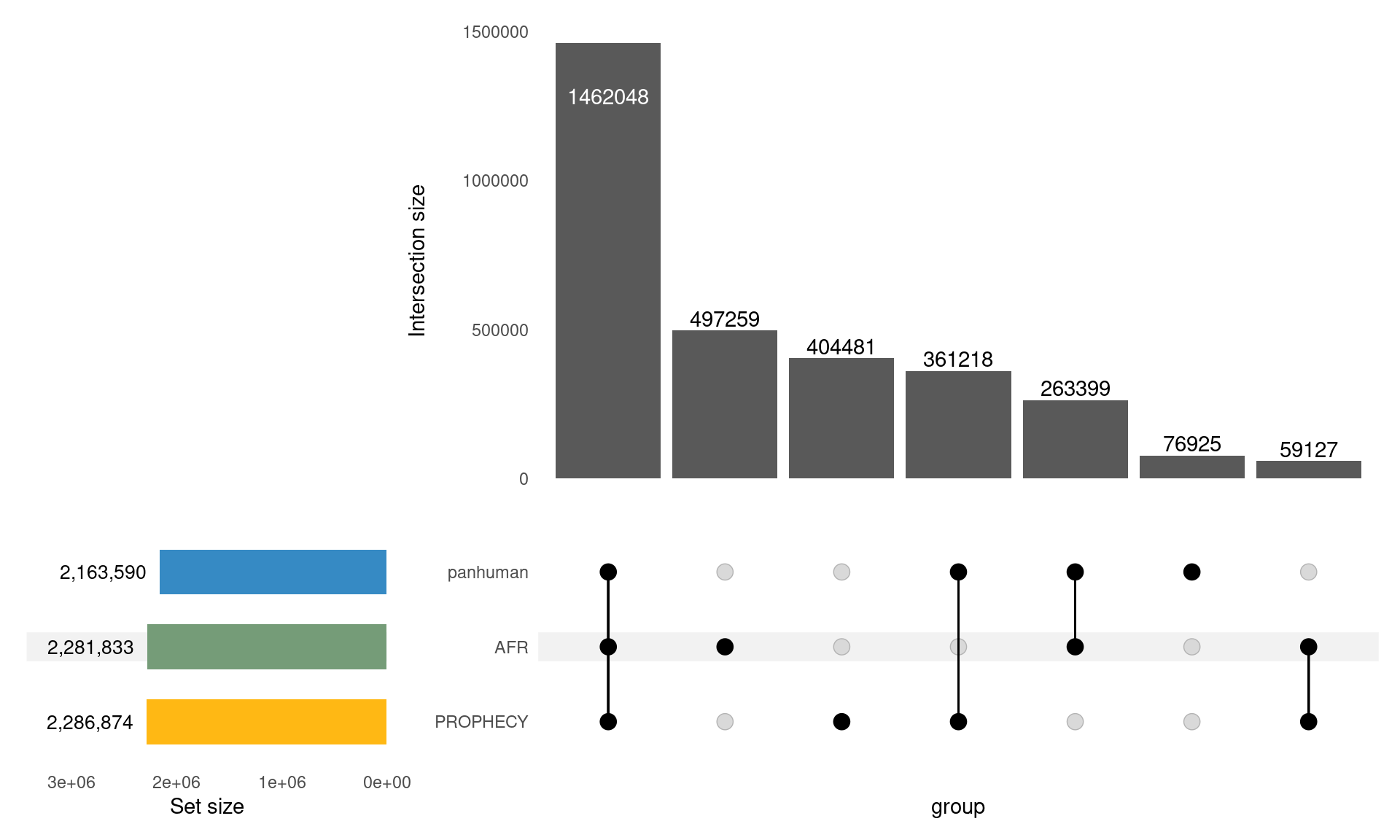
PROPHECY Variants Overlapping Exons
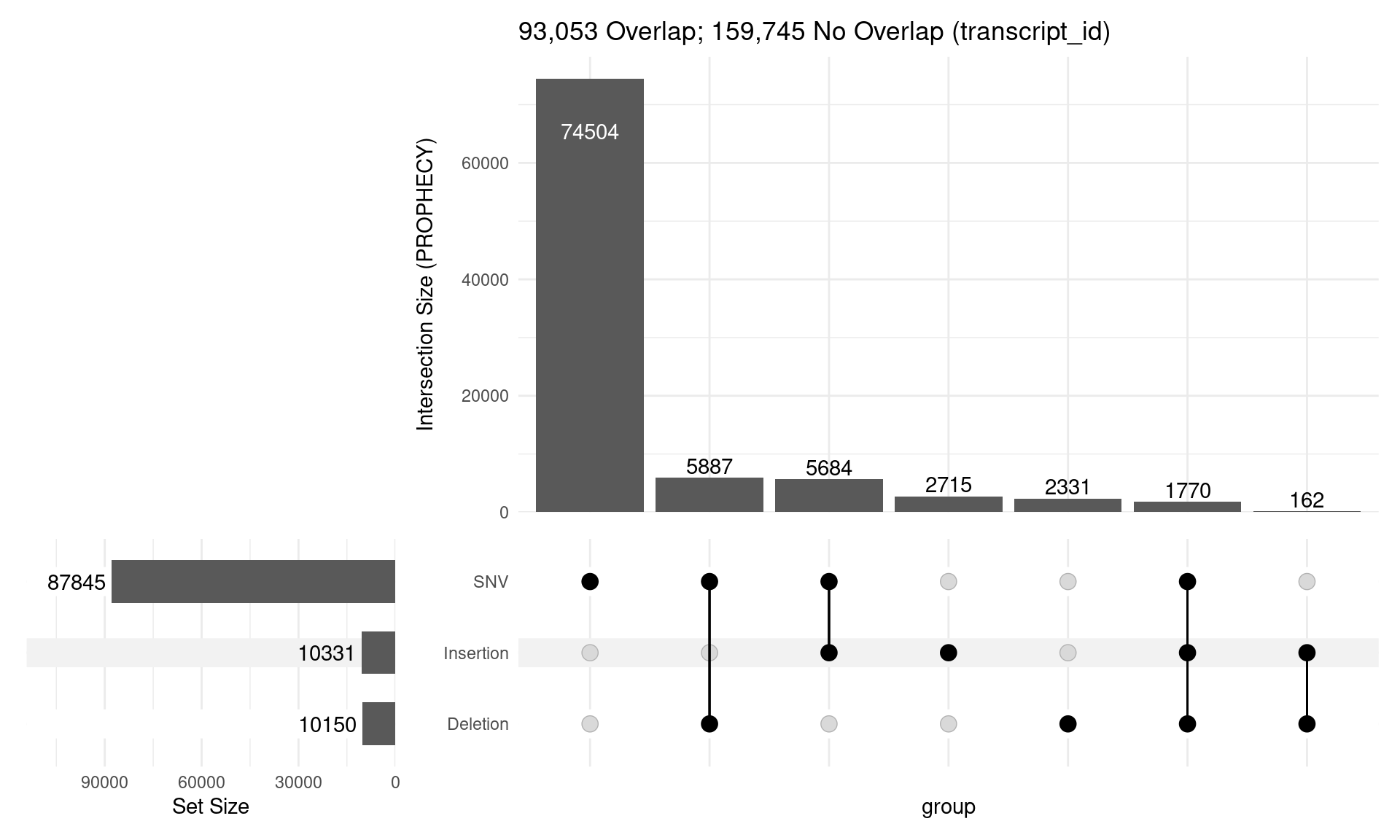
Key Questions
- Are technical metrics improved?
- Does it really make a difference?
- Does any difference actually capture the underlying biology?
- STAR is deterministic for given reference
- Change the reference even slightly \(\implies\) no guarantees
salmonis not deterministic
- Some of the changes will just be noise, but how much is noise?
Technical Analyses
- No real change to number of reads aligning (
STARorsalmon) - Very noticeable changes within alignments
- (Nearly) all technical metrics appear to be supportive of a variant modified reference
- First step is to check uniquely aligned reads from STARconsensus…
Uniquely Aligning Reads (STARconsensus)
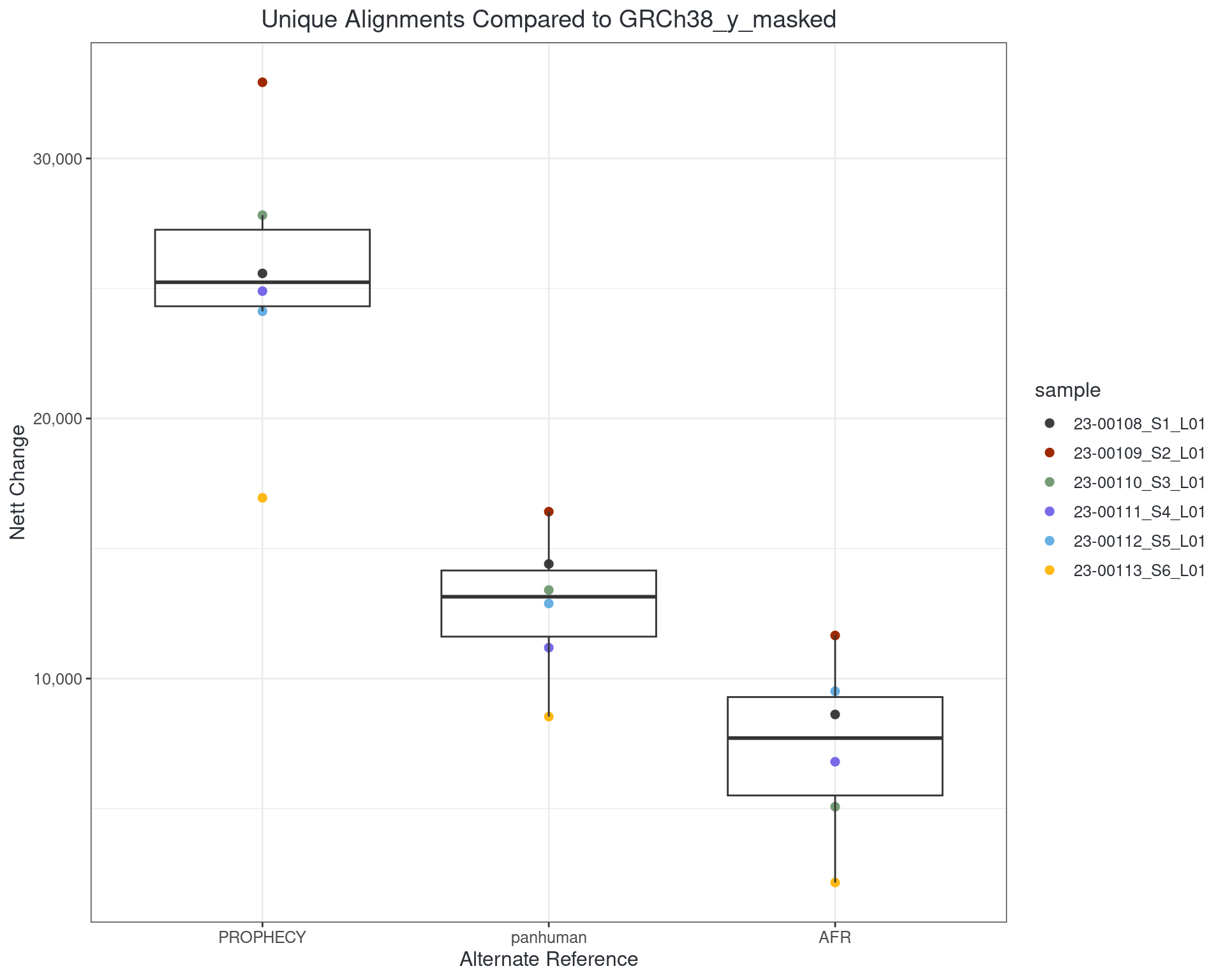
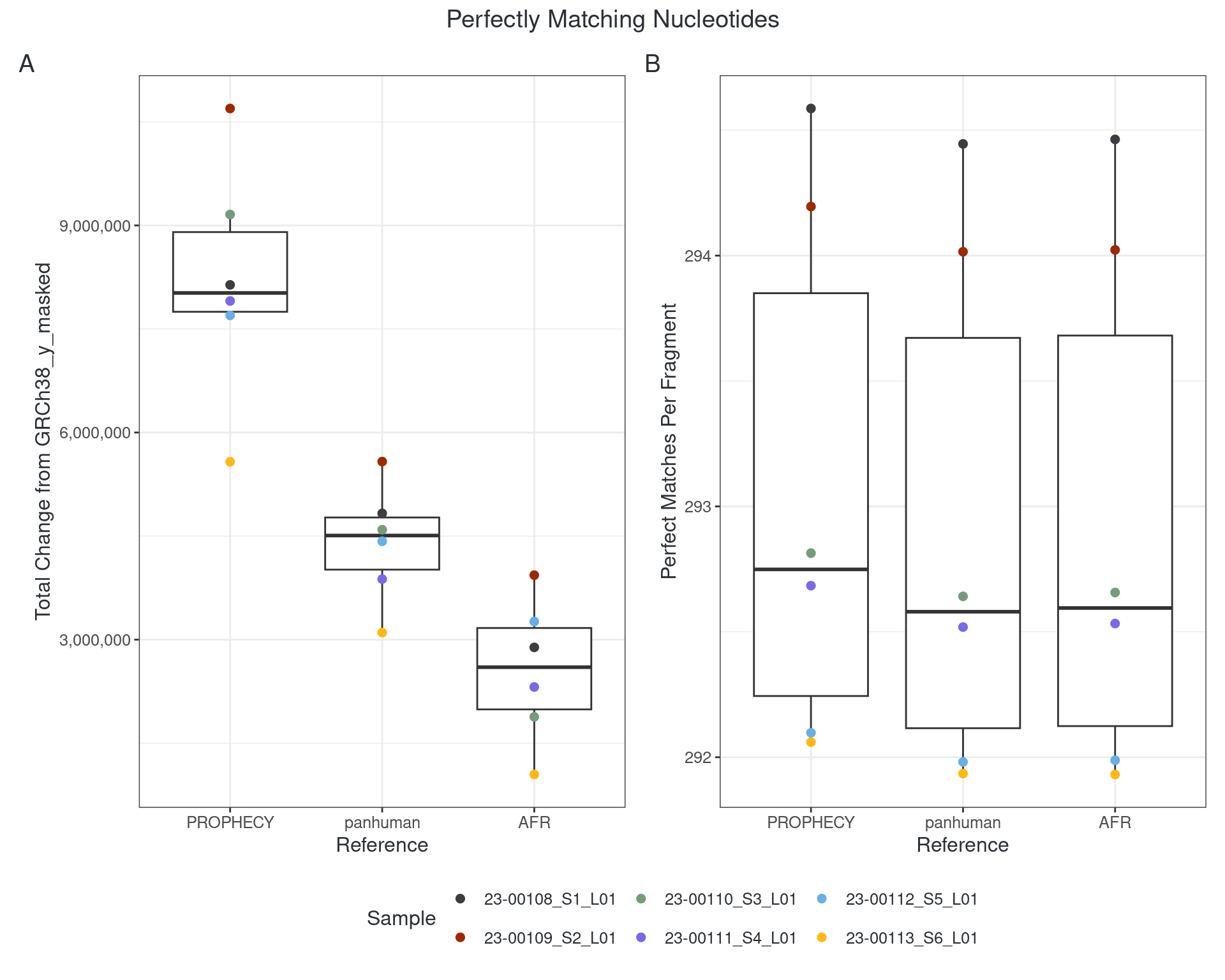
However…
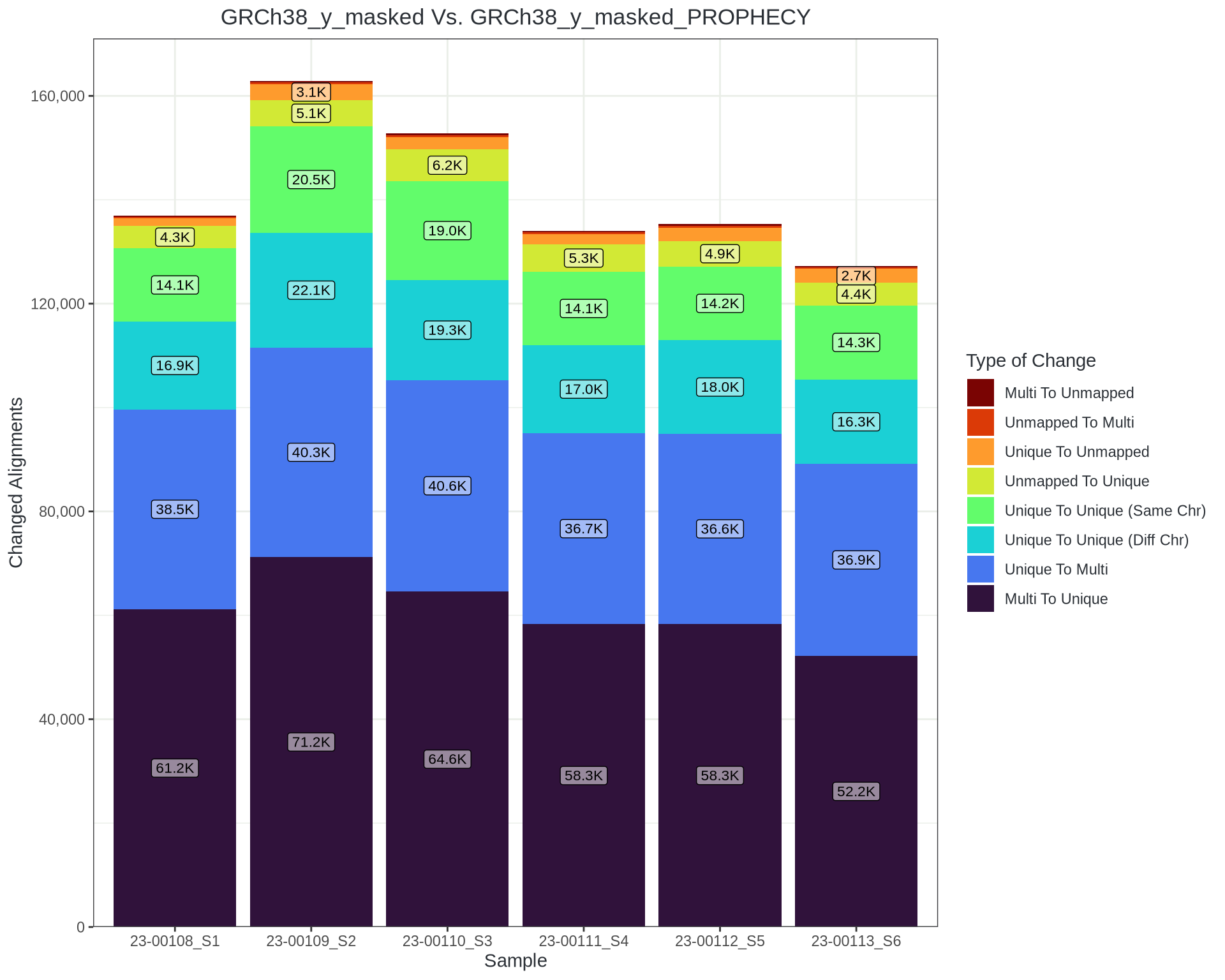
- About 1 in 300 reads moves
- About 30-40,000 reads remain uniquely aligned but in a different location
- How much is noise?
- Are the changes shared between samples?
- Do the changed reads overlap a variant?
- Where exactly are they moving from and to?
Reads Overlapping Variants
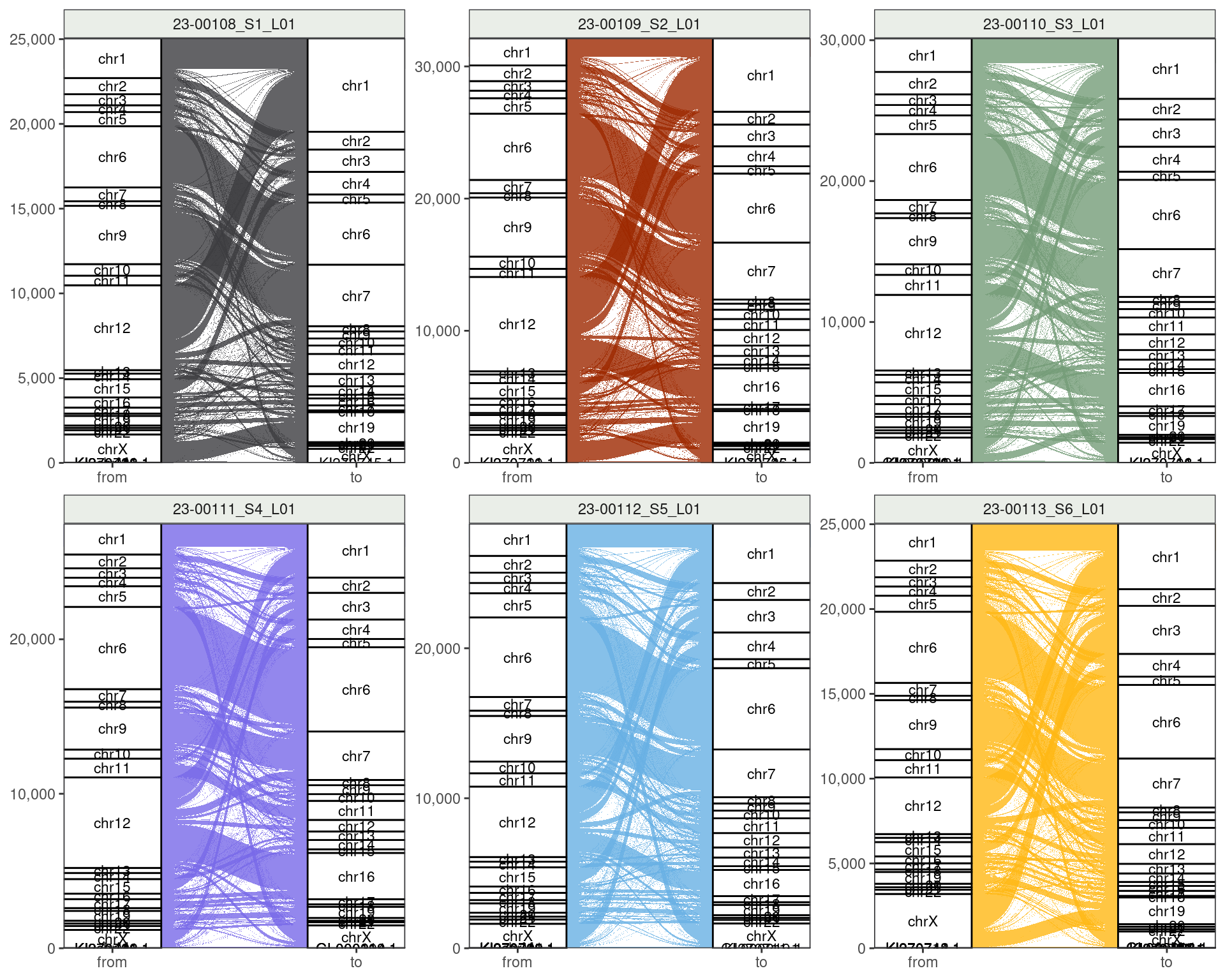
Taking reads where gapped alignments overlap a variant
- chr12 to chr1 shared by all
- Also chr9 to chr7
- chrX to chr19 shared by 1, 2, 6
- chr12 to chr16 shared by 2 \(\rightarrow\) 5
Reads Overlapping Variants
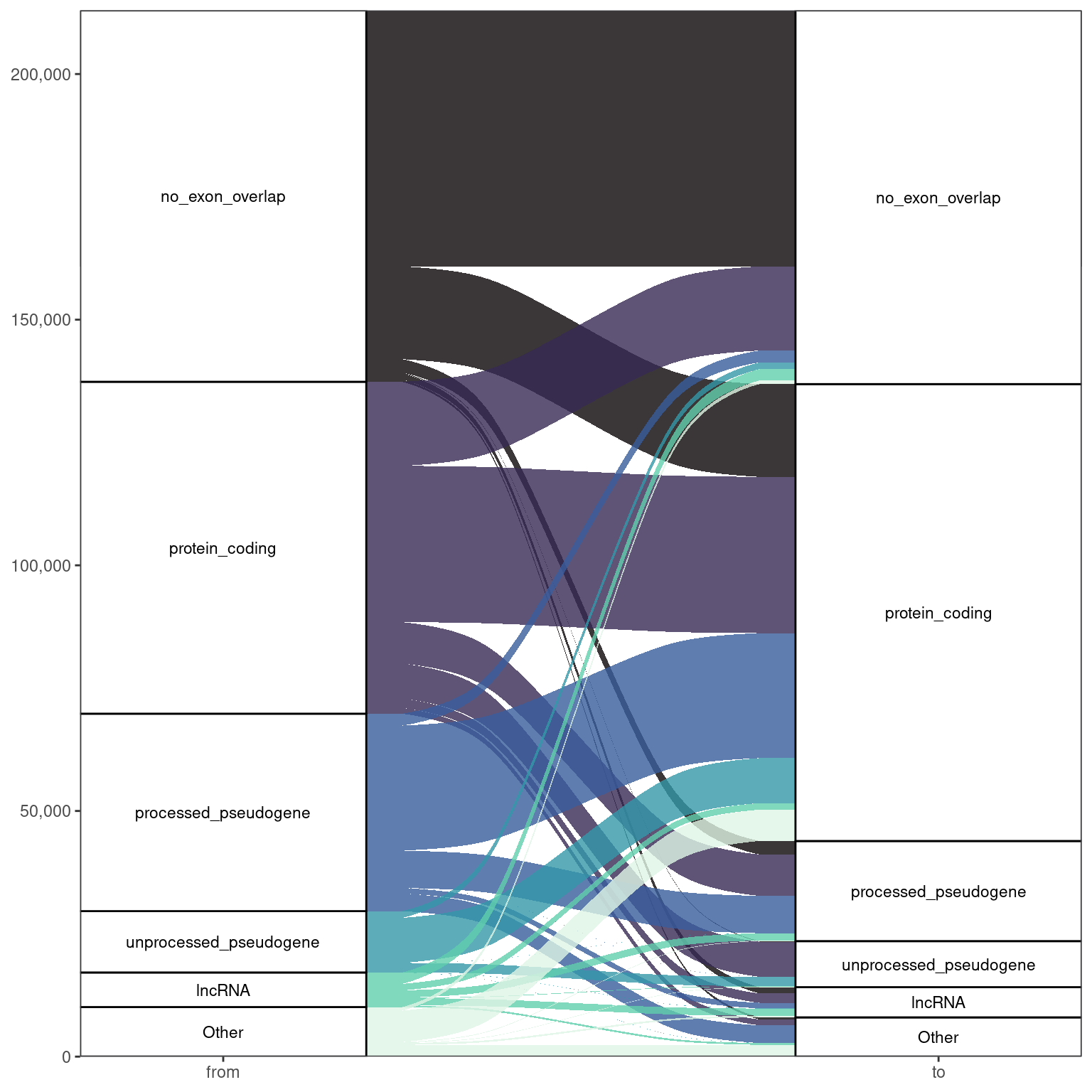
Taking all samples combined
- Increase in alignments to protein coding genes
- Decrease in pseudogenes
Common Genes Being Impacted
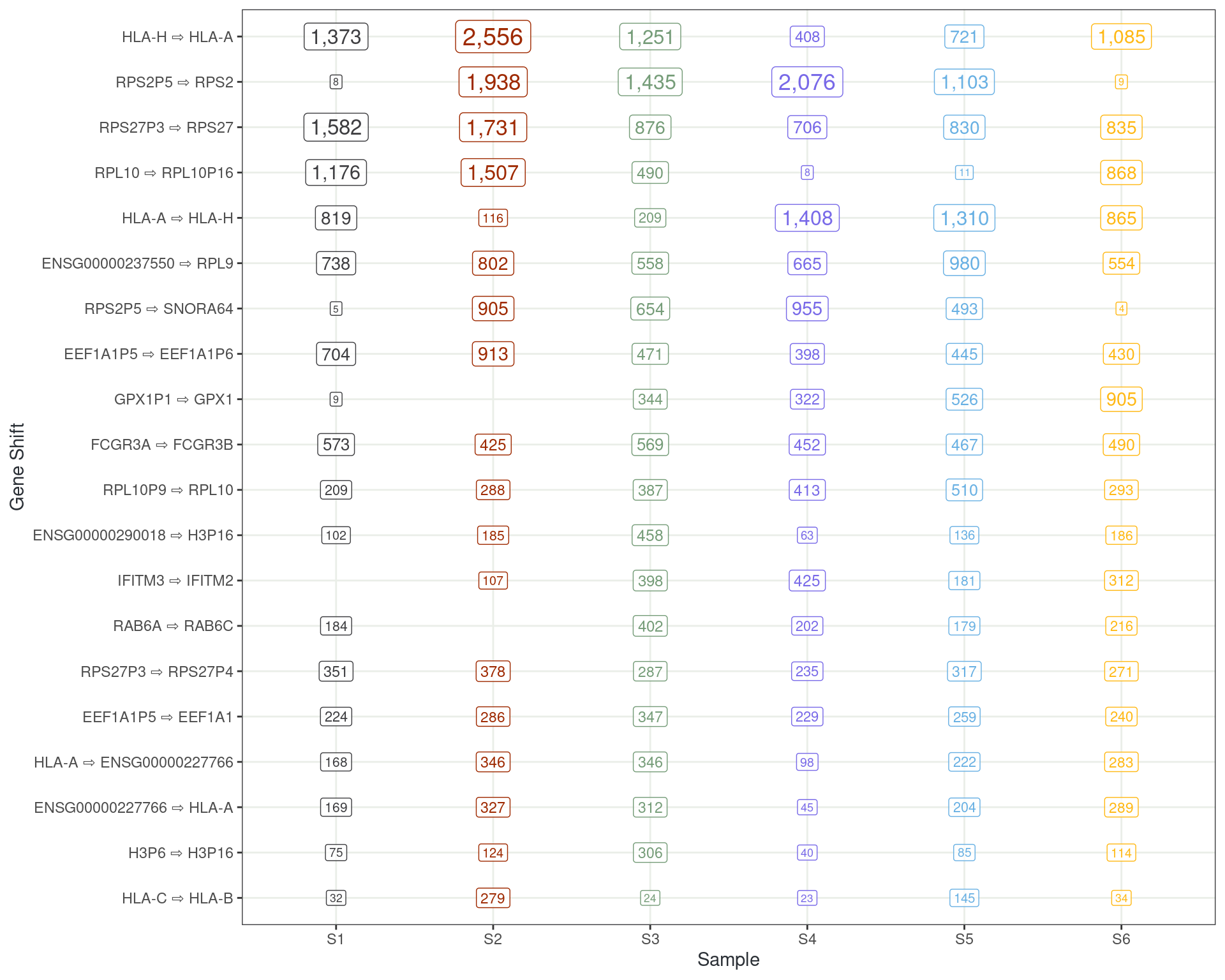
Taking the 20 most common shifts
- HLA-A
- HLA-H
- Ribosomal proteins
- Histone Proteins
Quick Summary
- These are only the uniquely-aligned reads from STARconsensus
- Don’t overlap alignment positions in either reference
- Haven’t looked at unmapped \(\rightarrow\) mapped, or multi-mappers
- Looks like it may be consistent & true biology?
- I got these results Wed night so I’m still processing them mentally
Differential Gene Expression Analysis
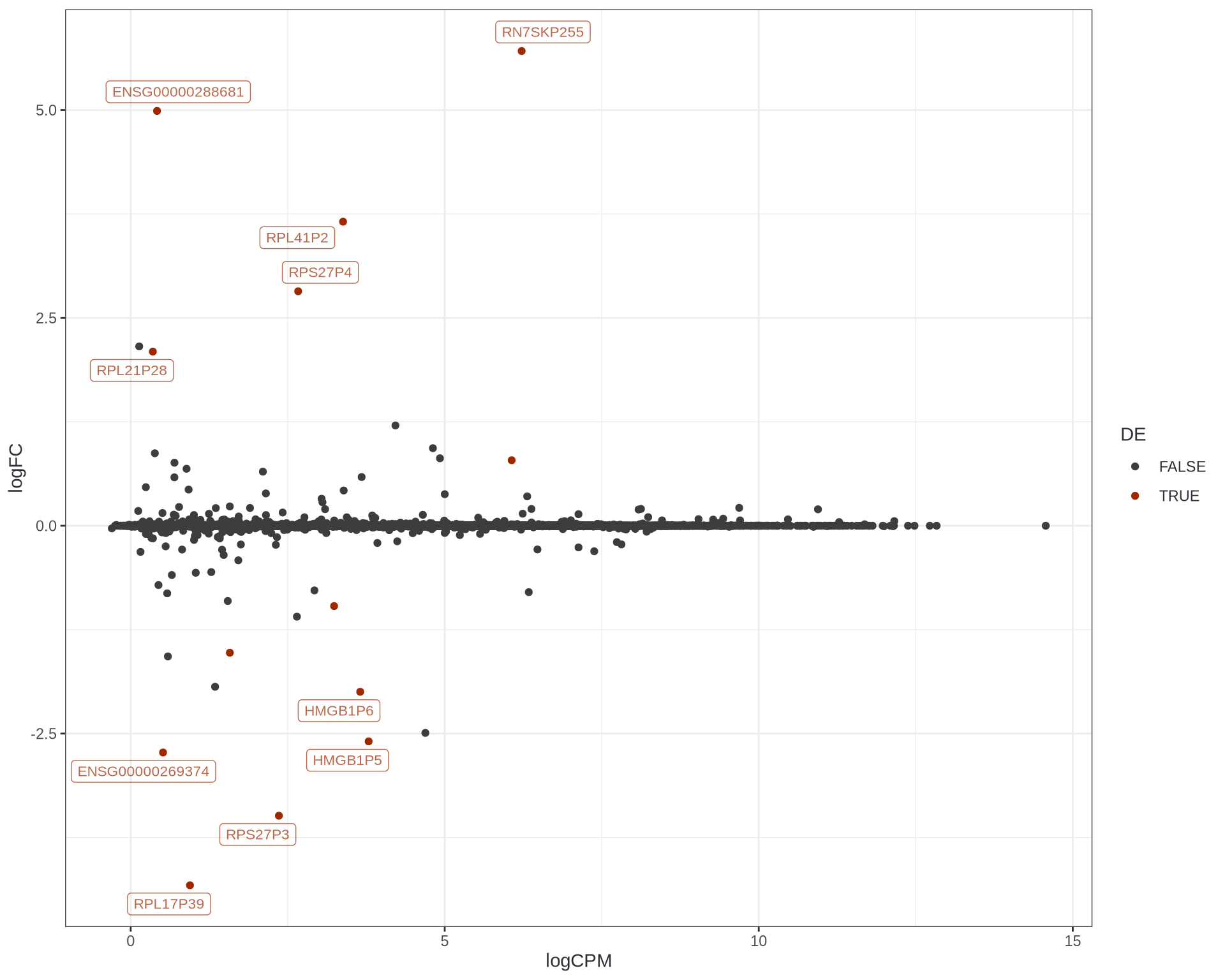
- GRCh38 vs PROPHECY-modified
- Identical input reads
- All changes are alignment induced
Genes Impacted Variably
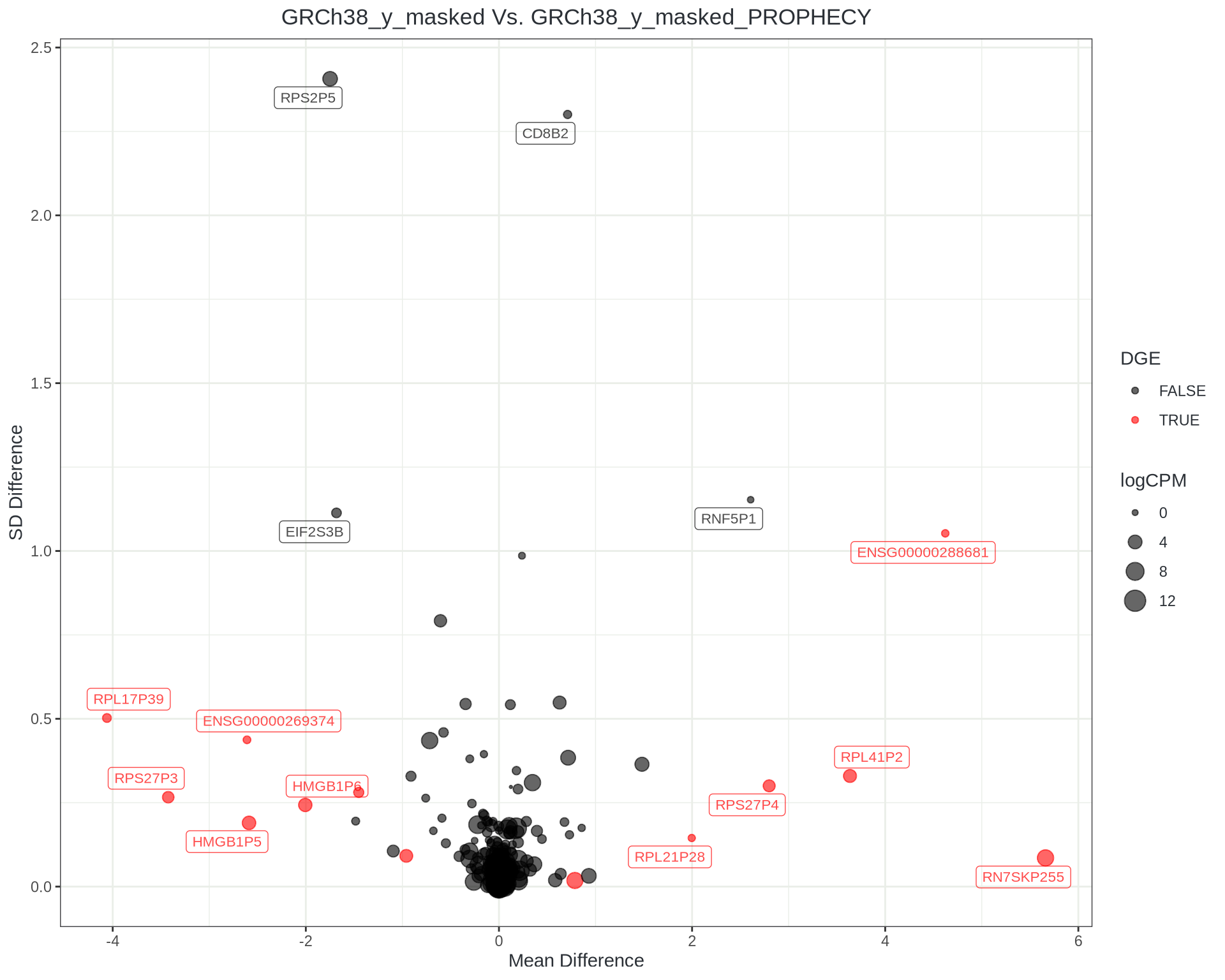
Taking expression estimates within each reference:
- Find the difference in counts per gene & sample
- Take mean & sd of differences
- Consistent changes will be low-variability, far from zero
- Inconsistent changes high-variability but near zero
- Like looking through an MA plot with variability as 3rd dimension
What about salmon + transmogR
Key Differences to STAR
- Only ~30% of reads align uniquely
- Largest NH:i tag is ~24,000
- 90% of reads align with NH:i < 10
- BAM files directly connect read ID to transcript
- Makes comparison much simpler
- How do the transcript-level counts change?
- How do the overdispersion estimates change?
Salmon Metrics
- Increase in Properly-Paired alignments (+3000)
- PROPHECY has strongest increase in correctly oriented fragments (+10,000 ISF)
- Decrease in umapped reads (-2000)
- Minor change in library sizes
- Similar numbers of detected transcripts (>1 count)
Overdispersion Estimates
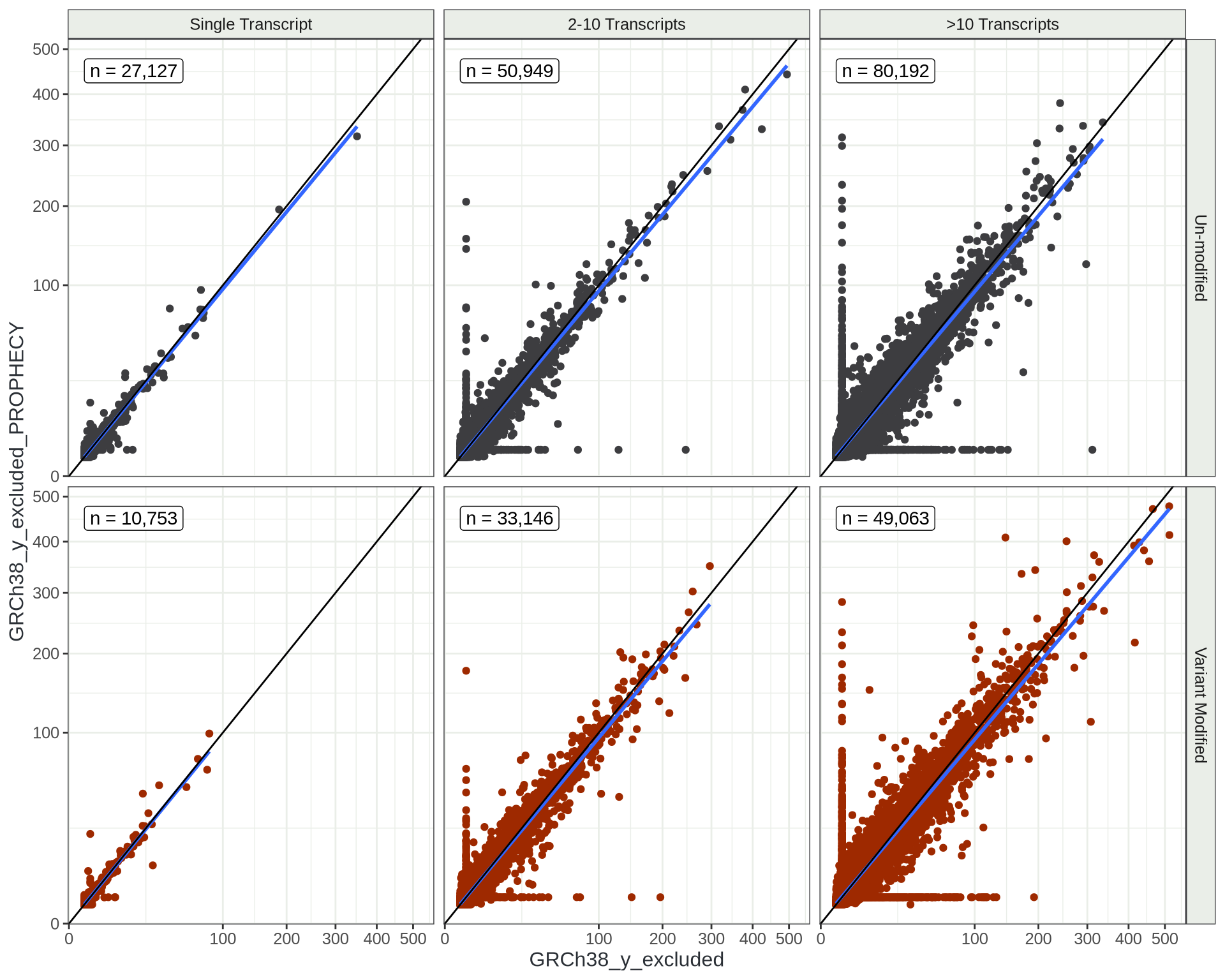
Looking at how the number of transcripts / gene also impacts these
Scaled Transcript-Level Counts
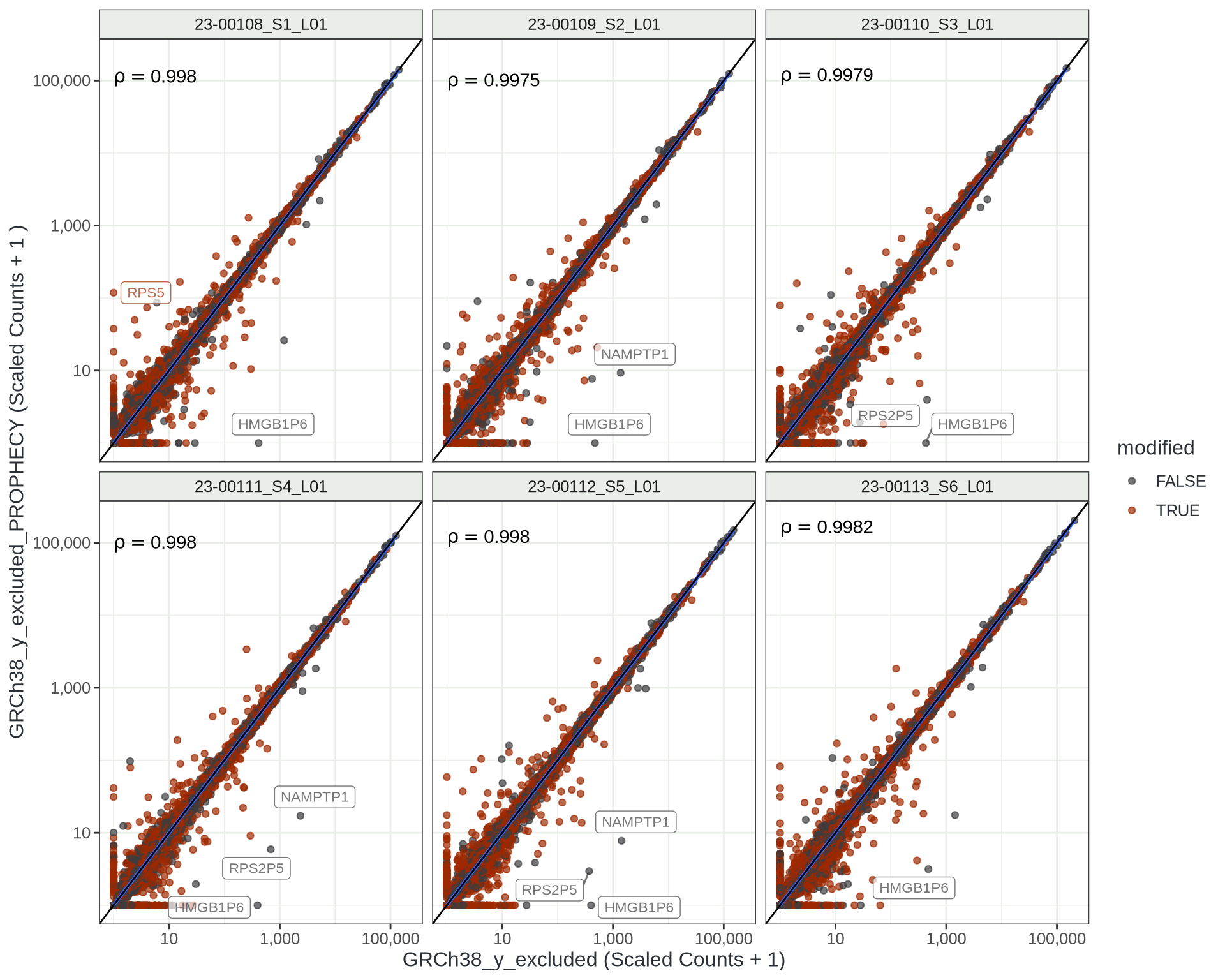
This seems quite acceptable
Variable Transcripts
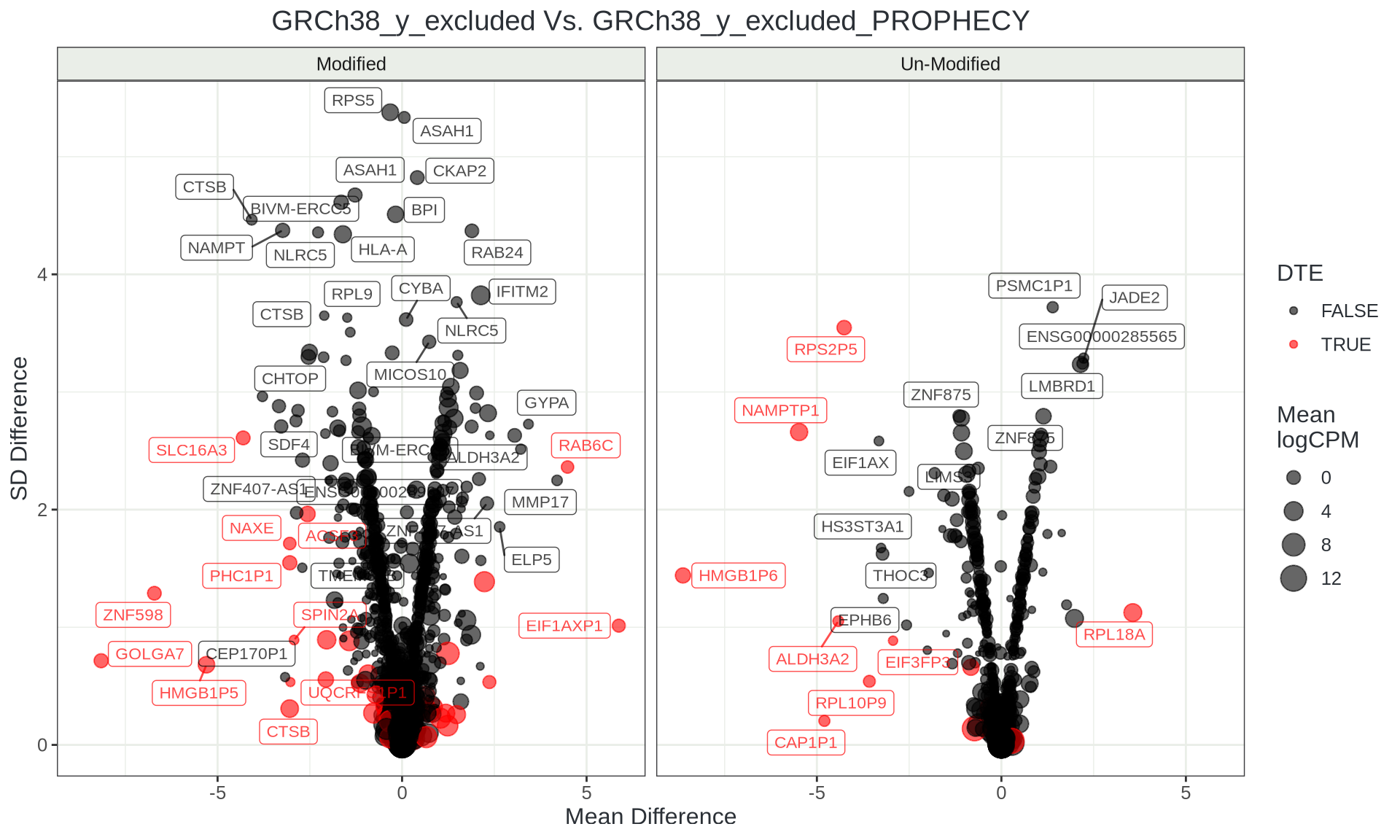
Tracking Individual Reads
- Can easily track a read & all it’s transcripts
- Subset data to reads with NH:i \(\leq 10\)
- Subset reads to those aligning to a modified transcript either before or after
- Unique-mappers are far less informative for transcript-level alignments
- Haven’t checked for moves between chromosomes
- Transcript & Gene-Level moves are more fine-grained
Tracking Reads Which Change Transcripts
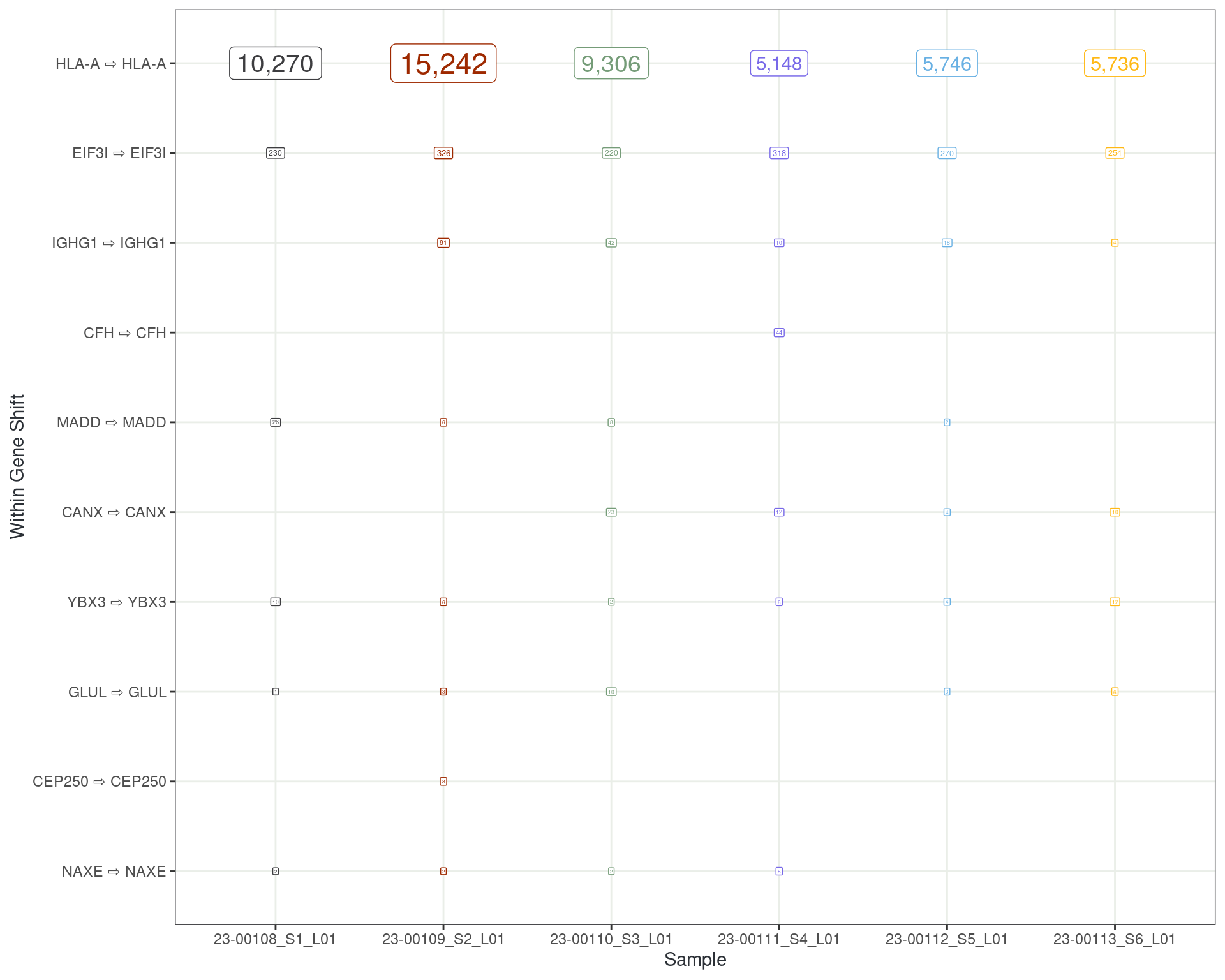
Reads changing transcript within a gene
Tracking Reads Which Change Transcripts
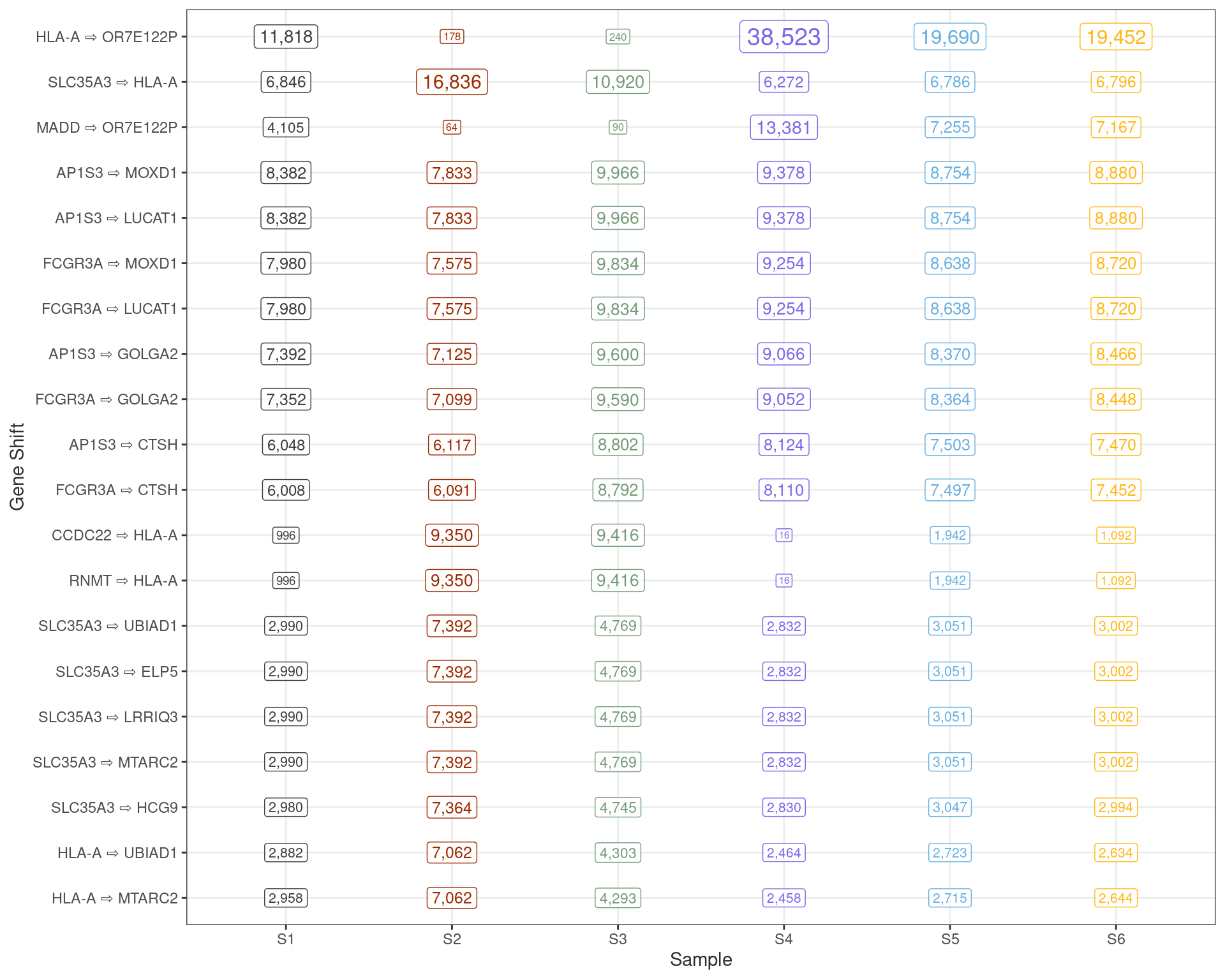
Reads changing transcript to different genes
Changing Transcripts: Gene-Level Categories
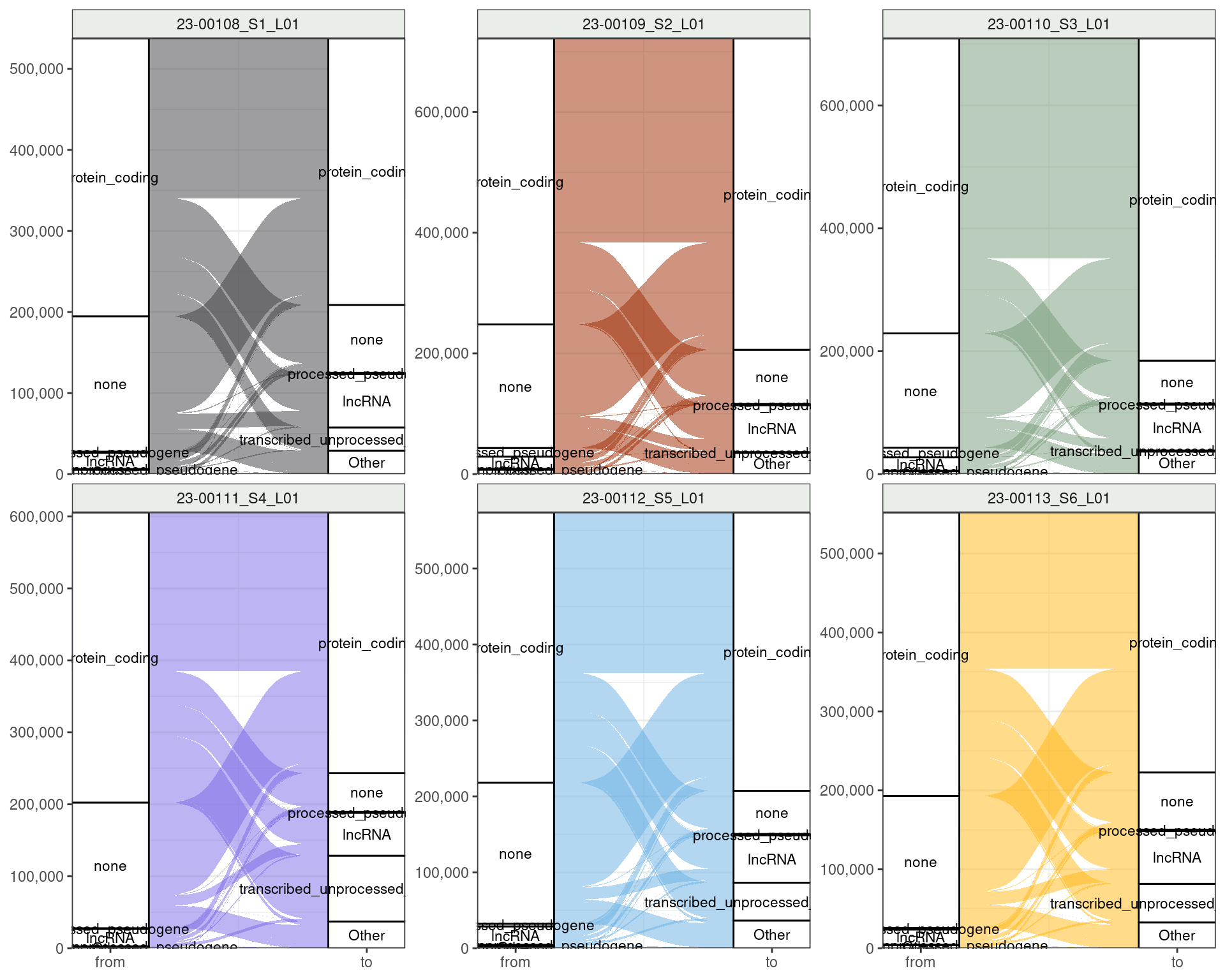

Shows a nice decrease in reads mapping nowhere
Changing Transcripts: Transcript-Level Categories
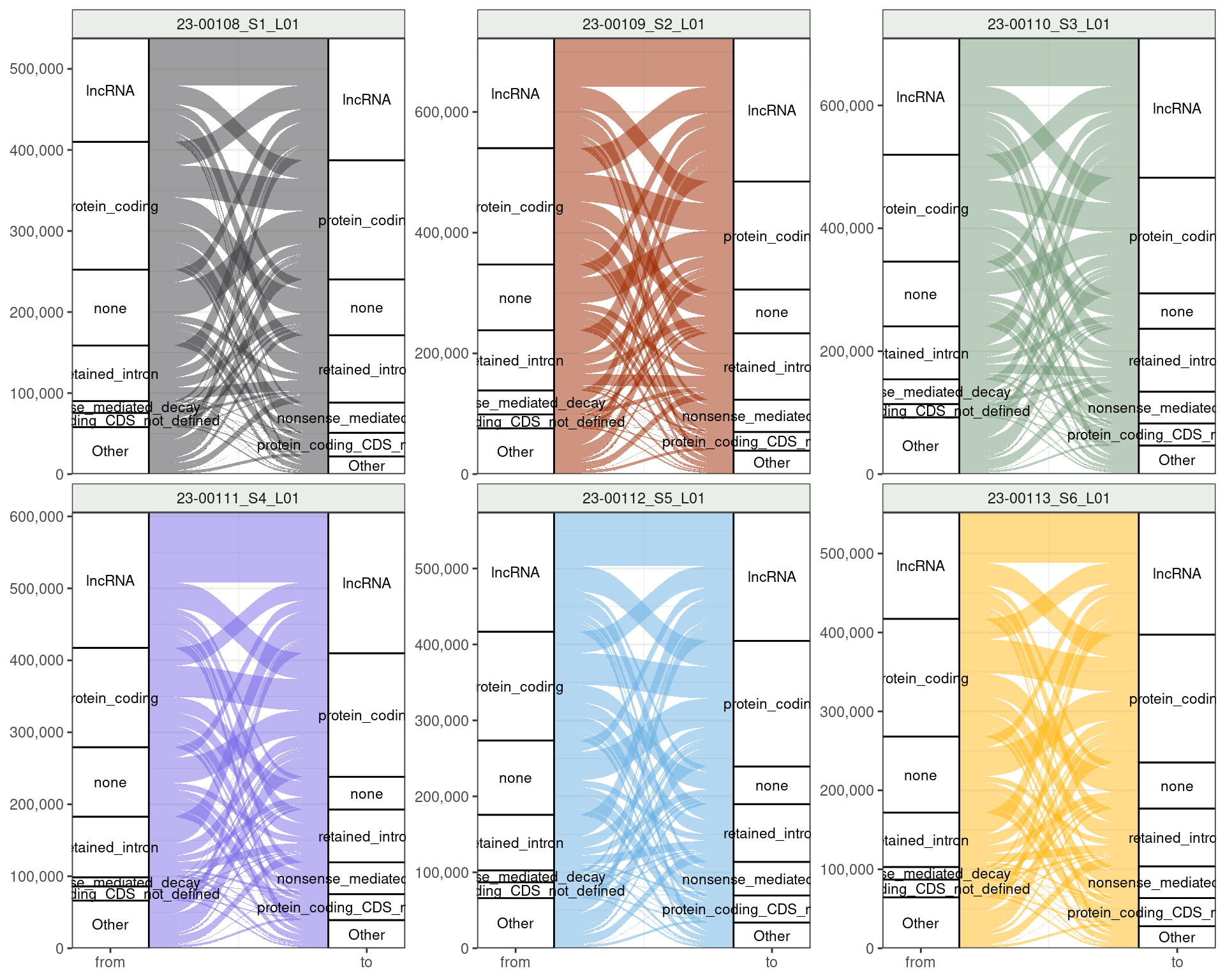
Lots of change between but no categories really sing out
Where To From Here
- Personalised Variant sets will give us the best estimate of ground-truth
- Will run STARconsensus in Diploid mode
- Haven’t figured out how to do this for
transmogR+salmon - Will just choose randomly from Het sites (???)
- Running a mini 90-sample cohort to see how it really plays at scale
- Previously analysed by Prof Sam El-Osta’s team (Monash)
- Have a specific subset of complications
- How will WGCNA modules be impacted?
- How will true DGE / DTE results be impacted?
Acknowledgements
Black Ochre Data Labs
Alex Brown
Jimmy Breen
Sam Buckberry
Yassine Souilmi
Bastien Llamas
Katharine Browne
Liza Kretzschmar
Alastair Ludington
Holly Massacci
Sam Godwin
Kaashifah Bruce
Rebecca Simpson
Sarah Munns
Ashlee Thomson
TKI / ALIGN
Johanna Barclay
Amanda Richards-Satour
Justine Clark
Rose Senesci
Analee Stearne
Louise Lyons
Dawn Lewis
Mary Brushe
Karrina DeMasi
Phoebe McColl
NCIG
Hardip Patel
SAHMRI
Tash Howard
Marlie Frank
SAGC
Sen Wang
Paul Wang
Renee Smith
University of Adelaide
Lachlan Baer
Monica Guilhaus
Wenjun (Nora) Liu
Megan Monaghan

References

