ALIGN Capacity Building
The Why, What and How of Transcriptomics
November 7, 2023
Acknowledgement of Country
I would like to acknowledge that I’m presenting today from Kaurna Country.
I acknowledge the deep feelings of attachment and relationship of the Kaurna people to their Place.
I also pay my respects to the cultural authority of Aboriginal and Torres Strait Islander peoples from other areas of Australia online today, and pay my respects to Elders past, present and emerging.
Who Am I?
- Oldest of the nerds in Black Ochre Data Labs but still an ECR
- Unintentionally took the “winding path” approach to my PhD (2008-2018)
- Undertaken in a T Cell / Autoimmunity Research Group
- Developed a novel Bayesian Statistical model for alternate transcript detection (using microarrays)
- Coordinator of the University of Adelaide Bioinformatics Hub (2014-2020)
- Analysed dozens of RNA-Seq datasets (+ more)
- Developed multiple training workshops and undergrad/postgraduate courses
- Massive R nerd (5 Bioconductor packages)
Who Am I?
- First discovered microarrays and R in 2002
- Most statistical understanding has been learned through this lens
- Very different to epidemiology, study design etc
- Primary training is in biology (e.g. genetics, biochemistry)
- Majority of experience in bulk tissue (i.e. not single-cell)
- Also spent two years researching ER-dependent Breast Cancers
- Looking at Transcription Factors and DNA state (ChIP-Seq)
- Data Integration (RNA-Seq, ChIP-Seq, HiC etc.)
Why Transcriptomics?
Why?
- Transcriptomics is essentially the analysis of RNA
- High-throughput viewpoint
- Provides insight into a highly dynamic component of biology
- Long history of identifying biomarkers and establishing mechanisms
- Is essentially a hypothesis-generating step
- Often need to test hypotheses using conventional experimental approaches
Why?
PROPHECY \(\implies\) Preventing Renal OPHtalmic Events in CommunitY
- Key focus is Type 2 Diabetes and complications (CVD and CKD)
- Multiple ’omics layers: DNA (Genomics), DNA-methylation (Epigenomics), protein (Proteomics), metabolites (Metabolomics) etc
- Whole blood \(\implies\) large immune component (“white blood cells”)
- Can we see markers of the different stages of Type 2 Diabetes
(i.e. complications developing)
- By integrating with other -omics layers:
- Can we refine our hypotheses into underlying mechanisms?
\(\implies\) clinical intervention
- Can we refine our hypotheses into underlying mechanisms?
How Does Biology Even Work?
DNA
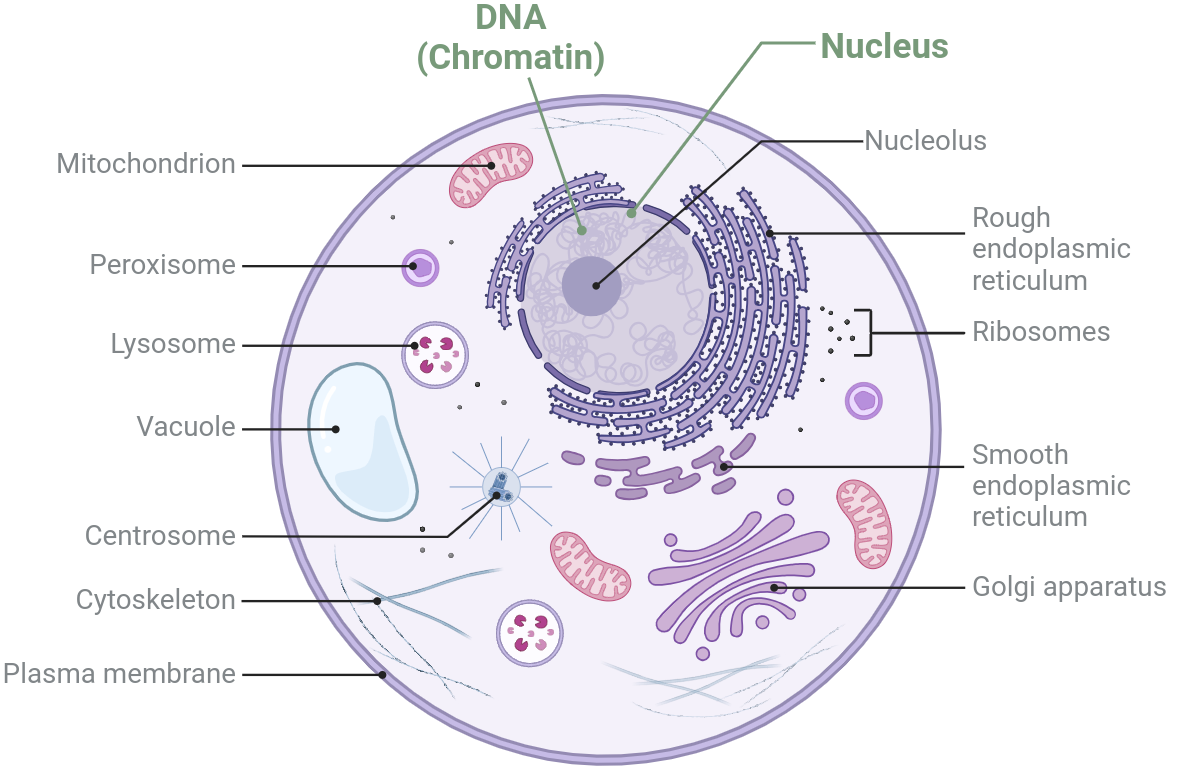
- Most are familiar with DNA and cells
- DNA stays in the nucleus
- Directly inherited from parents
\(\implies\) unique people - The same in every cell (roughly)
- Generally static (until cell division)
- Lots of interaction with other molecules: DNA-methylation, Histones etc.
DNA
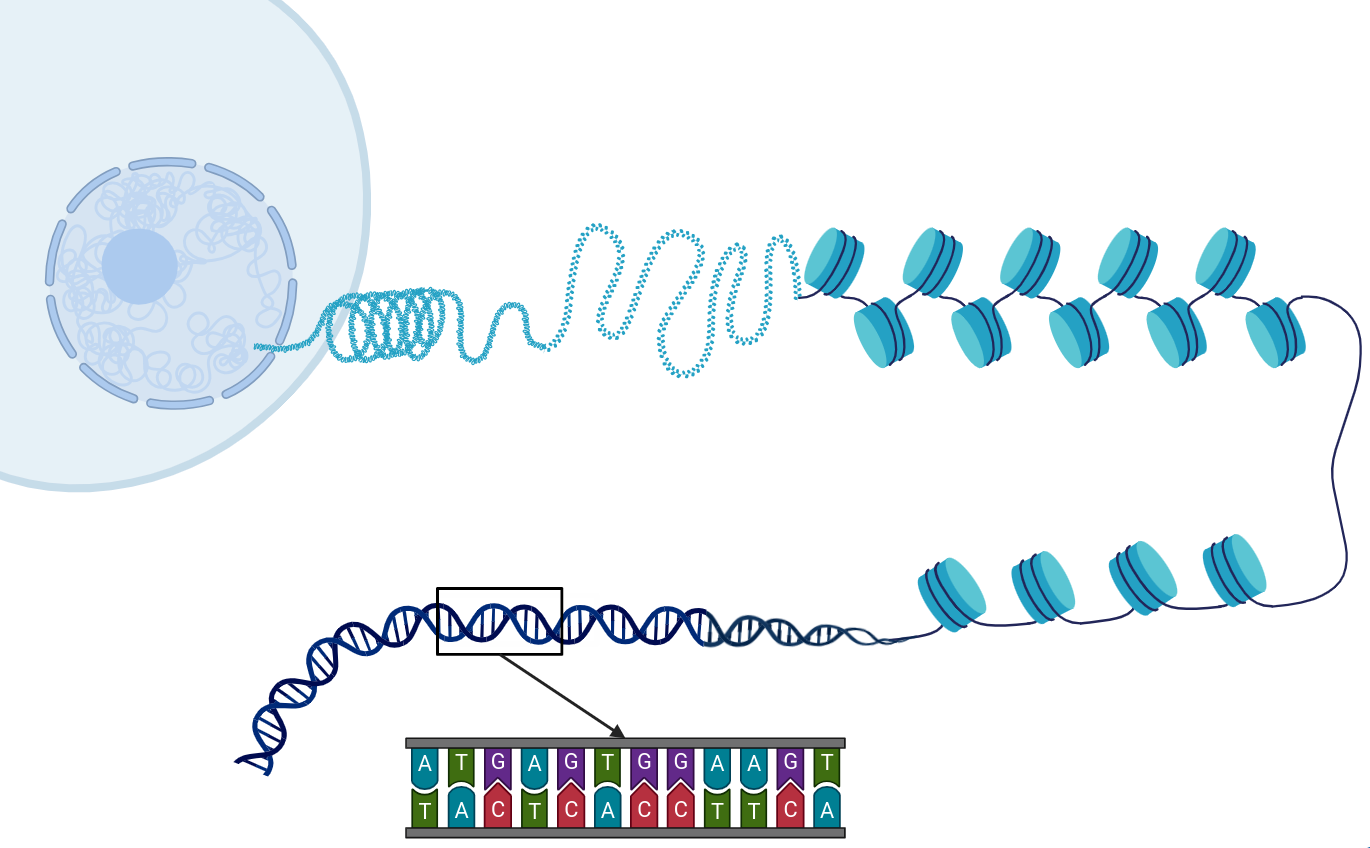
Each chromosome is a string of millions of nucleotides
- 4 nucleotides (or bases): A, C, G, T
- Double-stranded molecule, i.e. double helix
- Adds incredible stability
- Provides error handling mechanism
- Amazing amount of 3D structure & packaging
- Only forms the classic
![]() shape during cell division
shape during cell division
DNA
We often think of there being one unique, ideal reference sequence
- This is very far from reality
- Each copy of Chromosome 1 is different to the other copy within each of us
- Both copies are (essentially) the same between all of our cells
- No value judgements on similarity to reference \(\implies\) just who we are
- Same for all other chromosomes (1-22 + X)
- Scale this out to billions of people…
- Some regions are relatively invariant (needed for survival)
DNA
- Unique combinations of bases are ours and ours alone, for each of us
- Inherited from parents (+ a few spontaneous changes)
- We often refer to these unique sequences as variants
- Single nucleotides (SNVs)
- Missing or extra nucleotides (InDels)
- Larger structural variants (SVs) and repetitive elements (REs)
- Collections of variants within a population \(\implies\) genetic diversity
The Central Dogma of Biology
DNA is essentially a giant book of instructions
- Some regions are known as genes
- Transcribed into RNA
- Many RNA types (mRNA, miRNA, ncRNA, rRNA etc.)
- mRNA molecules are translated into proteins
- Groups of three bases encode a single amino acid
- Proteins do stuff i.e. pretty much everything
- Keratin \(\implies\) Hair structure
- Haemoglobin \(\implies\) Oxygen transport
- Antibodies \(\implies\) Detect “alien” molecules
Proteins
Proteins are the molecular machines that run our bodies
- Incredibly dynamic system
- Some are stable \(\implies\) some degrade rapidly
- Some exist only within cells \(\implies\) some are exported/imported
- Each cell-type (e.g. neuron, skin cell) will make a different set of proteins
- Some structural proteins may be common between unrelated cell-types
- Proteins can be heavily modified
- Phosphorylation, Sumoylation, Ubiquitination etc.
- Can be single molecules or bind to multiple partners
- Different functions or activities for every state
Proteins
Proteomics is the study of proteins
- Is very limited technique
- Tens of Thousands exist
- Can usually only identify hundreds (maybe 1-2000) from a sample
- We don’t understand much of the complexity
- Single modifications can have huge consequences
- Compared to DNA/RNA sequencing is a very immature field
- Technology is extremely variable and rapidly developing
What is Transcripomics?
Transcriptomics
Transcriptomics is the study of RNA molecules within a cell or tissue
- Not all genes are transcribed (i.e. expressed) in every cell
- ~60,000 annotated genes in the genome
- ~10-15,000 detectable genes in a given cell-type
- Transcriptomes are cell-type specific
- Related cell-types will often have great similarity, e.g. T-cell sub-types
- Often take a cell-line (or cell-type) and expose to a treatment
- Low variability \(\implies\) treatment response is easy(ish) to spot
Transcriptomics
- Transcriptomics is often an abundance-focussed analysis
- Represents a stable state of a cell
- Differential Gene Expression \(\implies\) abundance changes in response to stimuli
- Traditionally focussed on mRNA
- Common early assumption \(\implies\) mRNA abundance reflects protein levels
- Is sometimes true …
Transcriptomics
Our classic viewpoint is DNA \(\implies\) mRNA \(\implies\) Protein
Many other types of RNA play key roles
- miRNA bind to and degrade target mRNA
- lncRNA form highly complex structures
\(\implies\)can silence chromosomal regions (e.g. XIST)1
- rRNA are key components of ribosomes
- tRNA interact with ribosomes
\(\implies\) translate mRNA into proteins2

What is an RNA transcript?
- Known transcribed regions are defined as genes
- Transcribed from DNA \(\implies\) start to finish
- From the complete RNA transcript:
- Exons form mature transcript
- Introns are spliced out
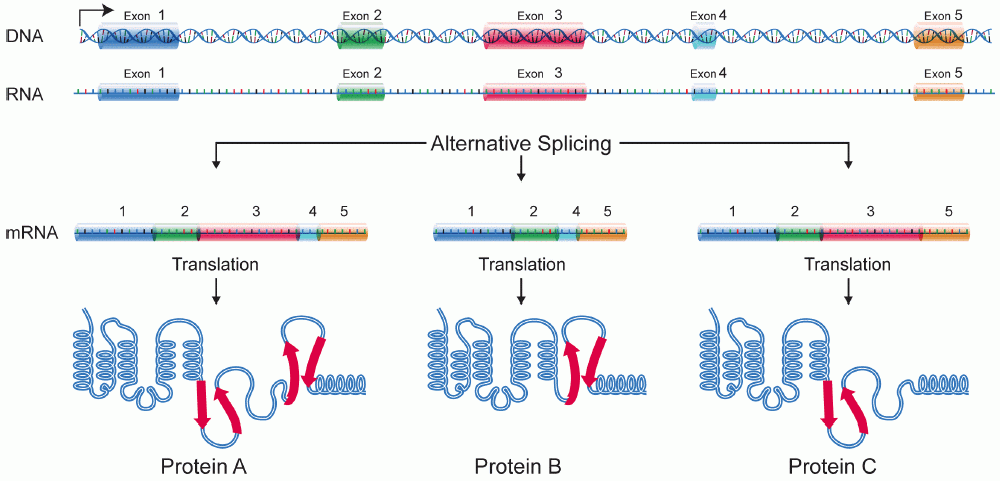
- ~38% of genes (~24,000) are transcribed into multiple transcripts
- Different transcripts \(\implies\) Different proteins (or lncRNA etc.)
How Do We Study Transcriptomics
Early Gene Expression Approaches
- The first high-throughput platform was Microarrays
- 3’ Arrays (Affymetrix)
- Gene-centric approach
- Could analyse 10-15,000 genes
- Abundance only analysis
- Many statistical tools developed in this context
limmahas been maintained by Gordon Smyth (WEHI) for >20years
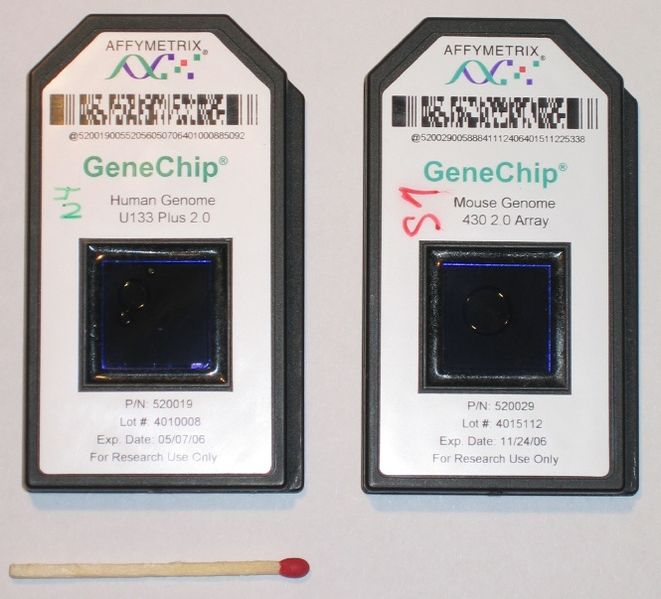

Microarrays
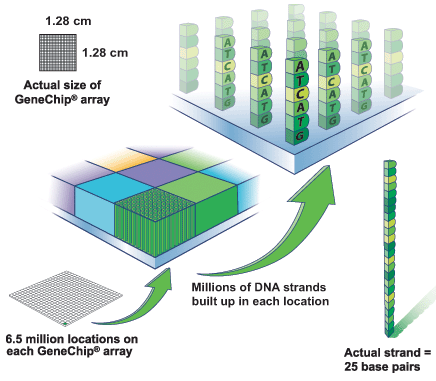
Known sequences at known locations
Labelled cDNA binds complementary target
- We analysed fluorescence NOT sequence data
Bulk RNA Sequencing
- Modern approaches involve sequencing the transcriptome
- Short Reads (Illumina) < 300bases: Quantitative
- Long Reads (Nanopore, PacBio); Semi-Quantitative
- Short reads still dominate
- Used in the PROPHECY transcriptomics layer
- Mature RNA transcripts may be short (8nt) or long (350,375nt)
Bulk RNA Sequencing
- All cells from each sample (or tissue) are lysed and RNA extracted
- RNA is fragmented (250-500bp)
- RNA Quality is assessed (RIN Score)
- RNA fragments are prepared for sequencing (library preparation)
- Converted to cDNA
- Add sequencing adapters
- PCR amplification
- Sequencing \(\implies\) ~30-50m reads/sample
How do we put this all back together and quantify?
Bulk RNA Sequencing
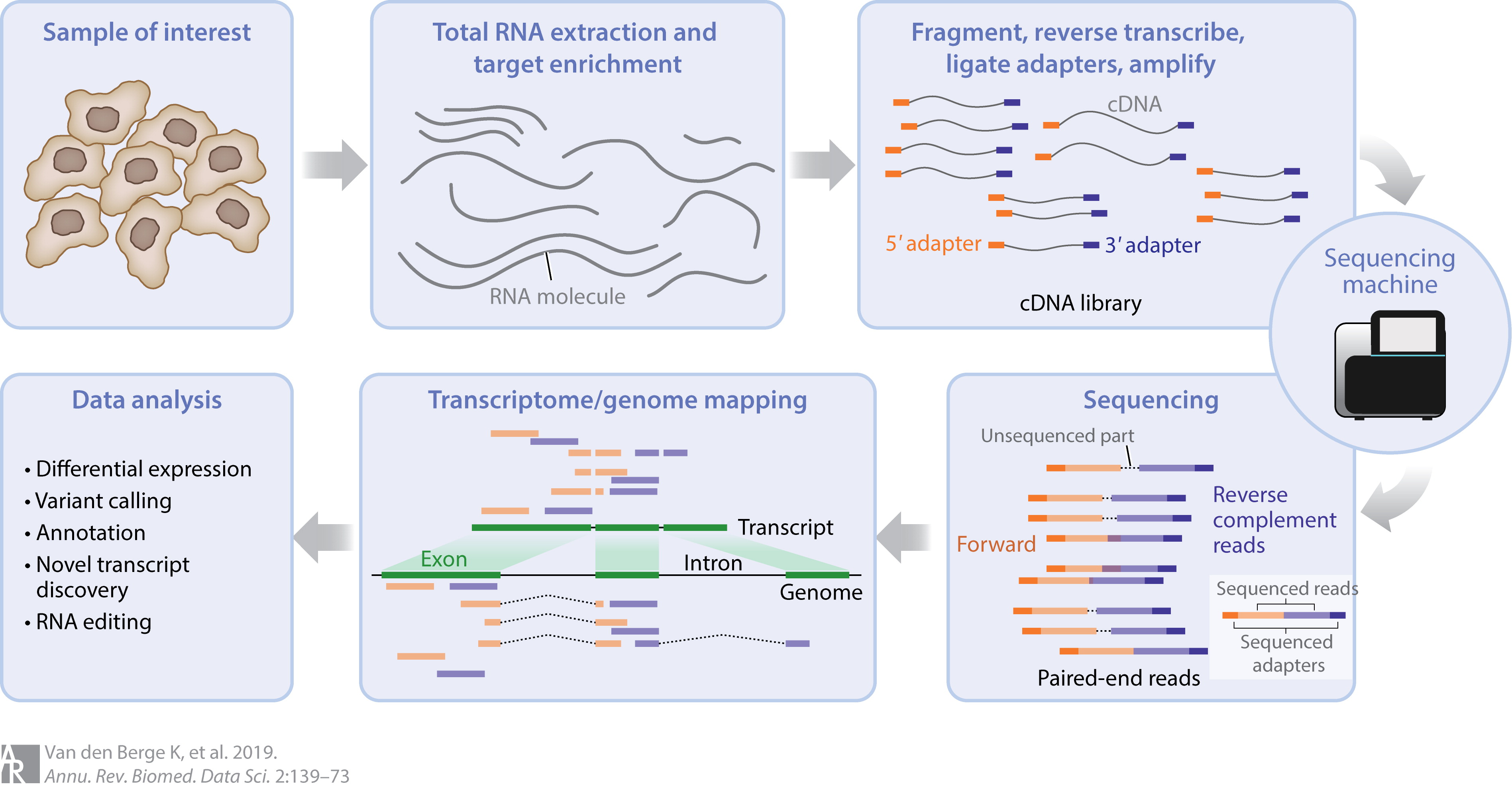
Two Approaches To Alignment
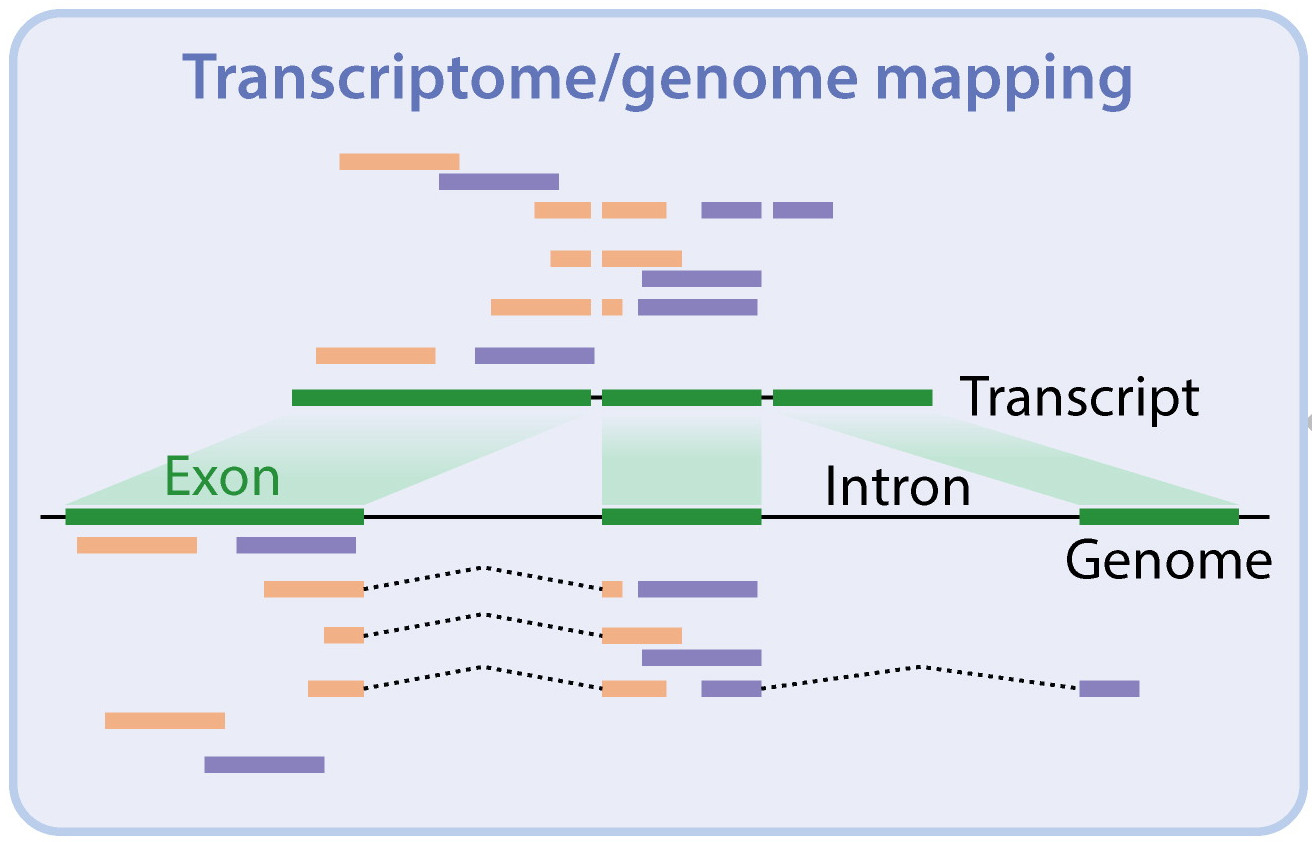
- Alignment to a reference genome
- Most align to one location \(\implies\) can see where ✓
- Brilliant for gene-level counts ✓
- Can call variants (allelic bias) ✓
- Not helpful for transcript-level ✗
- Alignment to a reference transcriptome
- Obtain transcript & gene-level counts ✓
- Many exons shared with multiple transcripts
\(\implies\) can model uncertainty ✓ - Don’t know where each read has aligned ✗
- No variant calling
Limitations
- Both approaches use 1 reference sequence \(\implies\) linear reference
- If 1 of each of our chromosomes matched exactly \(\implies\) the other copy won’t
- Current reference genome is GRCh38
- Transcripts are sequences derived from this
- Anchored to GRCh38 using co-ordinates
- Includes 517 extra sequences (scaffolds, patches, haplotypes)
- Do not exist in reality
- Some genes have transcripts defined on scaffolds not chromosomes
How Do We Manage Diversity?
Given that no real person matches the reference genome
- Can we improve the reference in general?
- Can we improve for a given study cohort
Two emerging approaches
- Modify a linear reference using representative variants
- Align to a reference graph which represents appropriate diversity
Modifying a Reference
- Tools exist for modifying a reference genome
- Unable to analyse at the transcript level \(\implies\) only gene-level
- BODL is developing the software to produce a variant-modified transcriptome
- Will enable counts at the transcript-level + uncertainty measures
- Current testing uses 1000 Genomes Project variants
- Shown to significantly improve mappings in other datasets
- Poorly representative of Australian Indigenous variation
Graph Based Approaches
- Telomere-to-telomere (T2T) assemblies contain no scaffolds
- Just chr1-22, X, Y & MT
- Individual assemblies \(\implies\) haplotype resolved
- Separate out both copies of chr1 etc.
- Can construct a reference graph
- Shared sequences between chromosome pairs are joined
- Variation represented as bubbles
- For >1 individual would be a pangenome graph
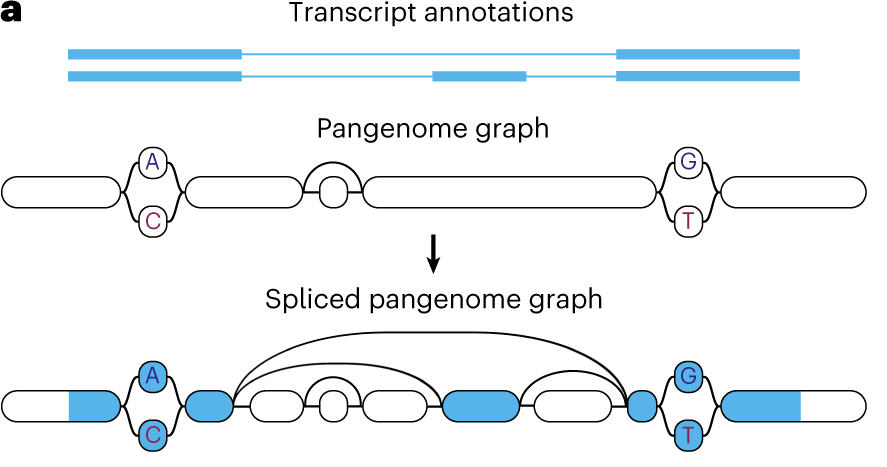
Graph Based Approaches
- A reference pan-genome graph is now available1
- Contains 47 haplotype resolved diploid T2T assemblies
- Representative of diversity?
- Explicitly align to the complete set of diversity (in the reference)
- Are the days of the linear reference dead? 😱
- How will this impact the PROPHECY transcriptomics layer?
- e.g. How do we integrate annotated regulatory elements from a linear reference
How Do We Make Key Discoveries?
Differential Gene Expression
Once we have counts (gene or transcript-level)
- Identify any technical issues (GC bias, failed samples etc.)
- Fit standard statistical models (GLM)
- Fairly simple in small Treatment vs Control comparisons
- Less straightforward with large, complex designs
- Sequencing generally done in batches of \(\leq\) 96 \(\implies\) batch effects
- Is identifiable technical noise which masks true biology
- Lead to ⇧ false discoveries ⇩ true discoveries
- Active area of development for large cohort studies (Terry Speed)
Differential Gene Expression


Network approaches
- DGE takes “significantly DE” genes and joins to try & form a story
- i.e. big changes in a few genes \(\implies\) biological consequences
- Network approaches look for larger shifts amongst correlated genes
- i.e. small changes across an entire pathway \(\implies\) biological consequences
- Far more flexibility with parameters \(\implies\) reproducibility?
- Recent research is improving this markedly
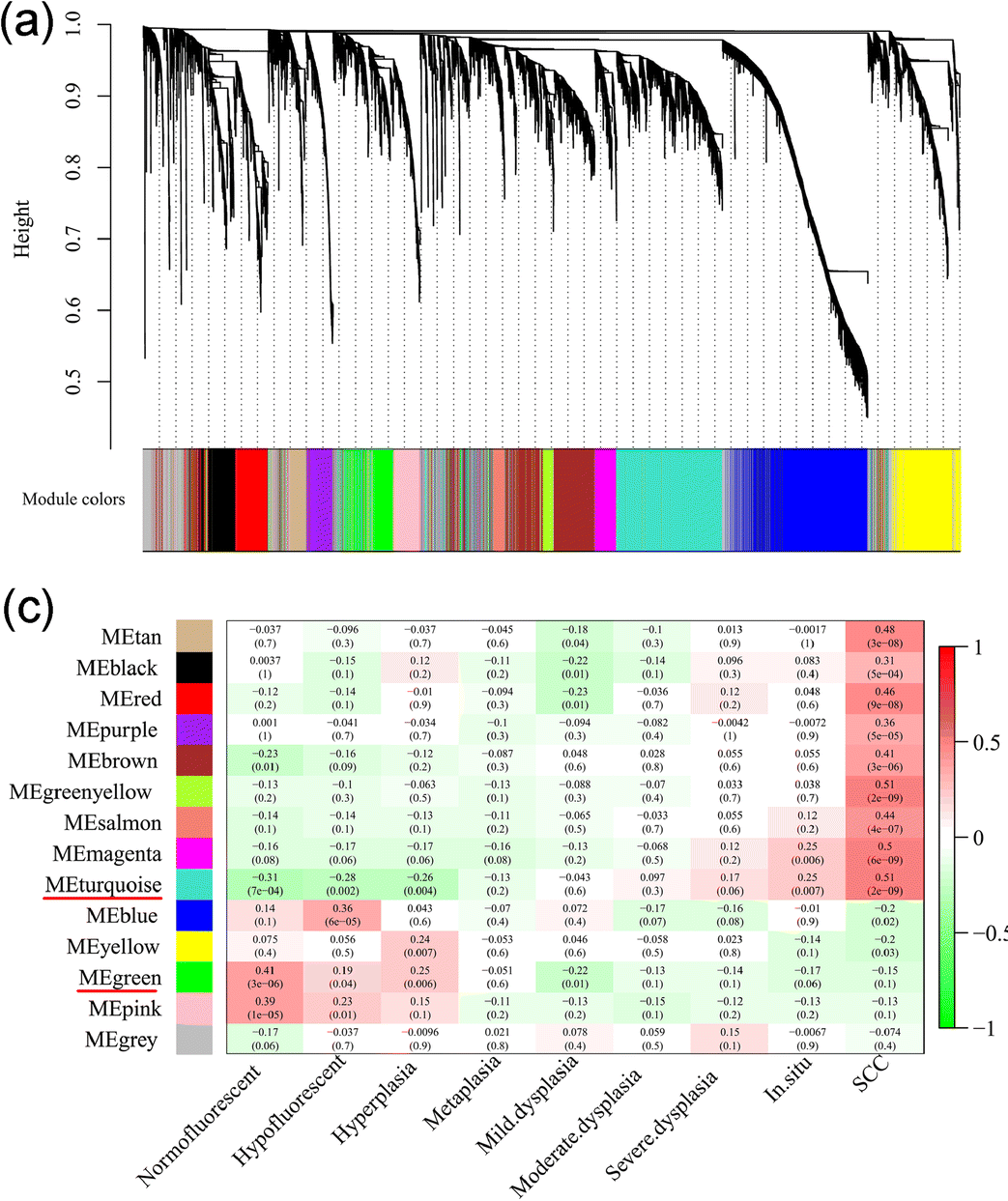
WGCNA
- Multiple approaches with WGCNA the biggest player
- Form correlation network
- Identify modules within correlation network
- Compare to predictor variables
- Identify underlying biology
Supervised Approaches
- Principal Component Analysis (PCA) is an un-supervised approach
- Components maximises variance
- Mainly used for QC & visualisation in transcriptomics
- Projection onto Latent Space (PLS) is a supervised approach
- Components maximise covariance with predictor variables
- Alternative approach to identifying groups of correlated genes
- No \(p\)-values 😱 \(\implies\) How can we ever publish?
eQTL and TWAS
- Expression Quantitative Trait Loci (eQTL)
- If RNA abundance is a quantitative trait \(\implies\) which variants are associated with this?
- Is there an association with phenotype (e.g. CVD development)
- Transcriptome-Wide Association Studies (TWAS)
- Integrates analysis eQTL with GWAS
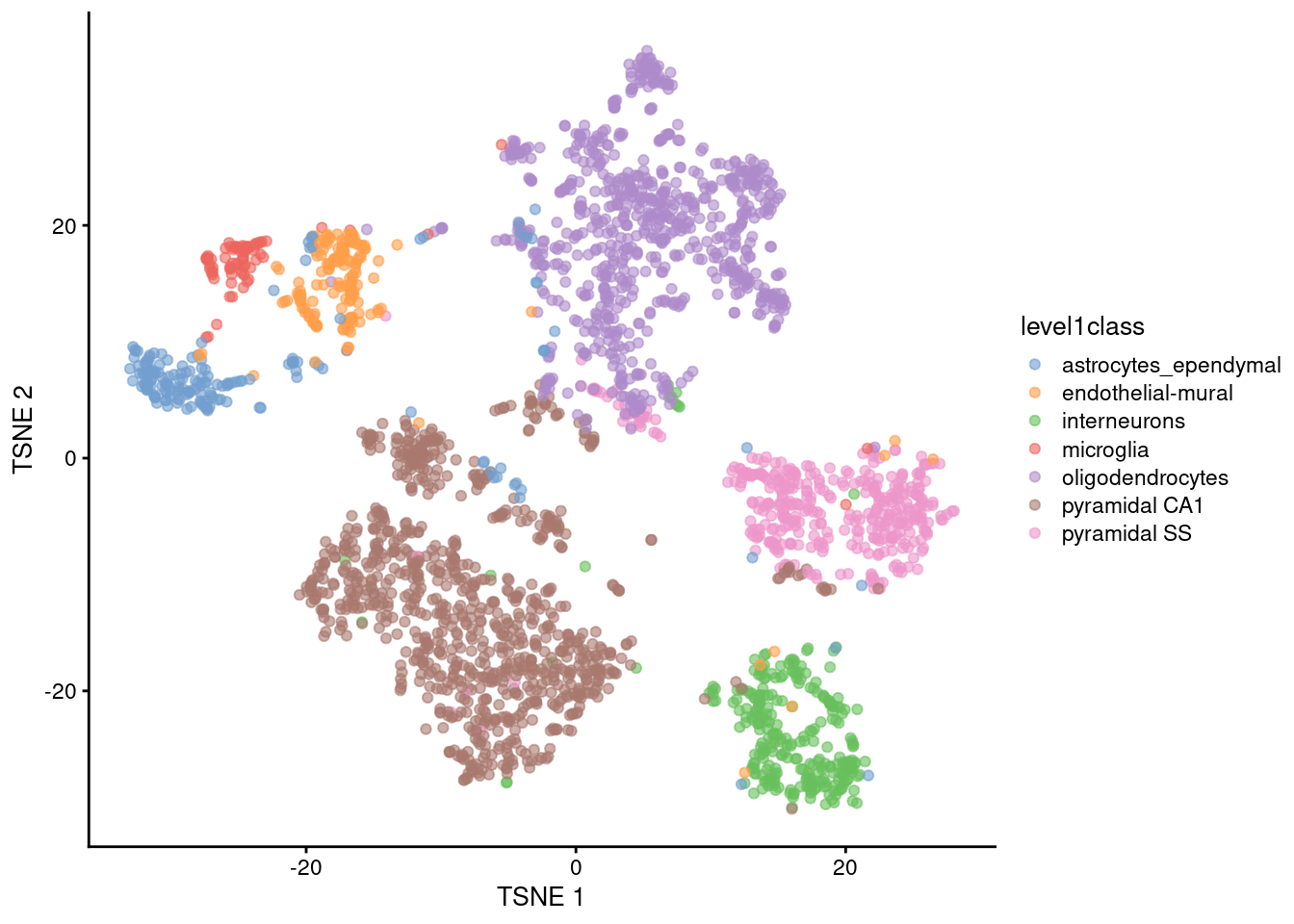
scRNA-Seq
- Retains the connection between transcript and cell-of-origin
- Huge numbers of ‘failure to detect’ expression (i.e. zero counts)
- Uses clustering to identify cell types within a sample
- Pseudo-bulk clusters for DGE analysis
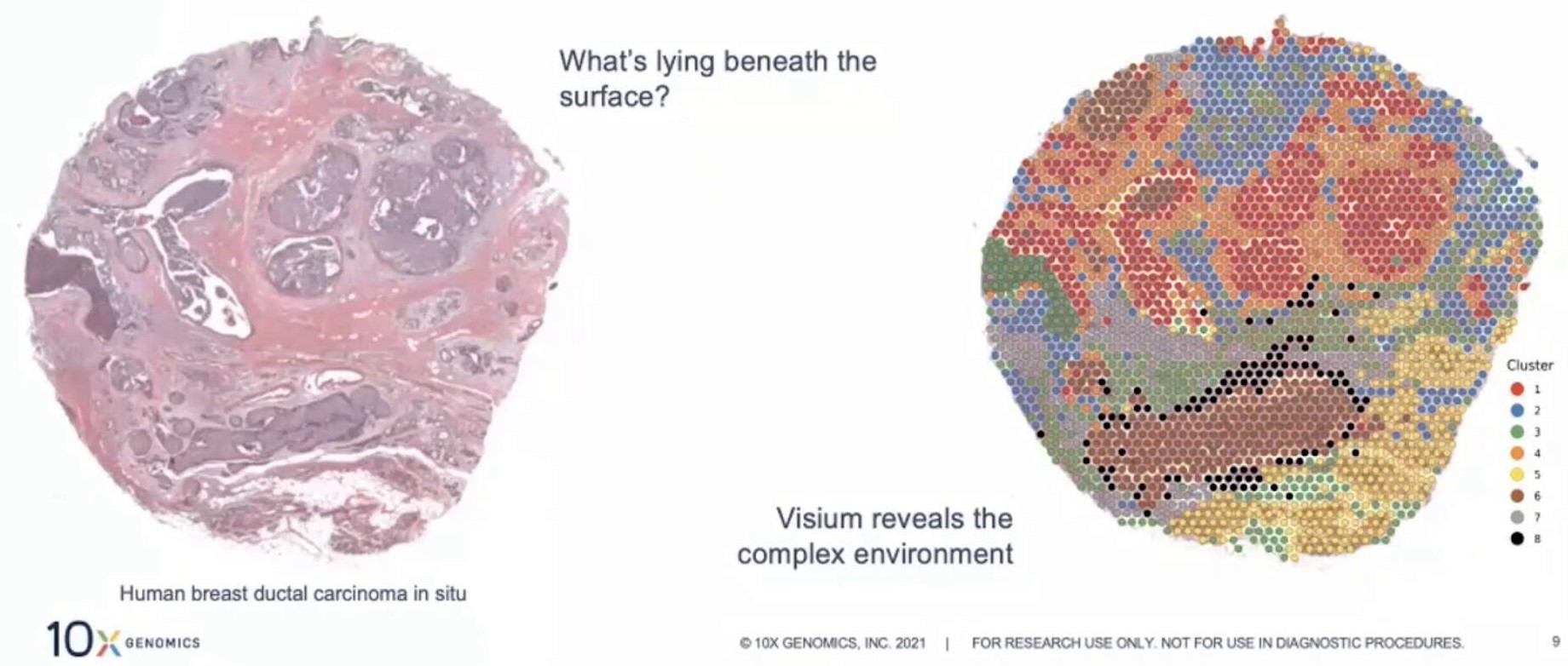
Spatial Transcriptomics
- Cells are held in place \(\implies\) transcripts identified within a region
- The current hot area in transcriptomics \(\implies\) Nature Methods “Method of the Year, 2020”
- Single-cell resolution is arguably here
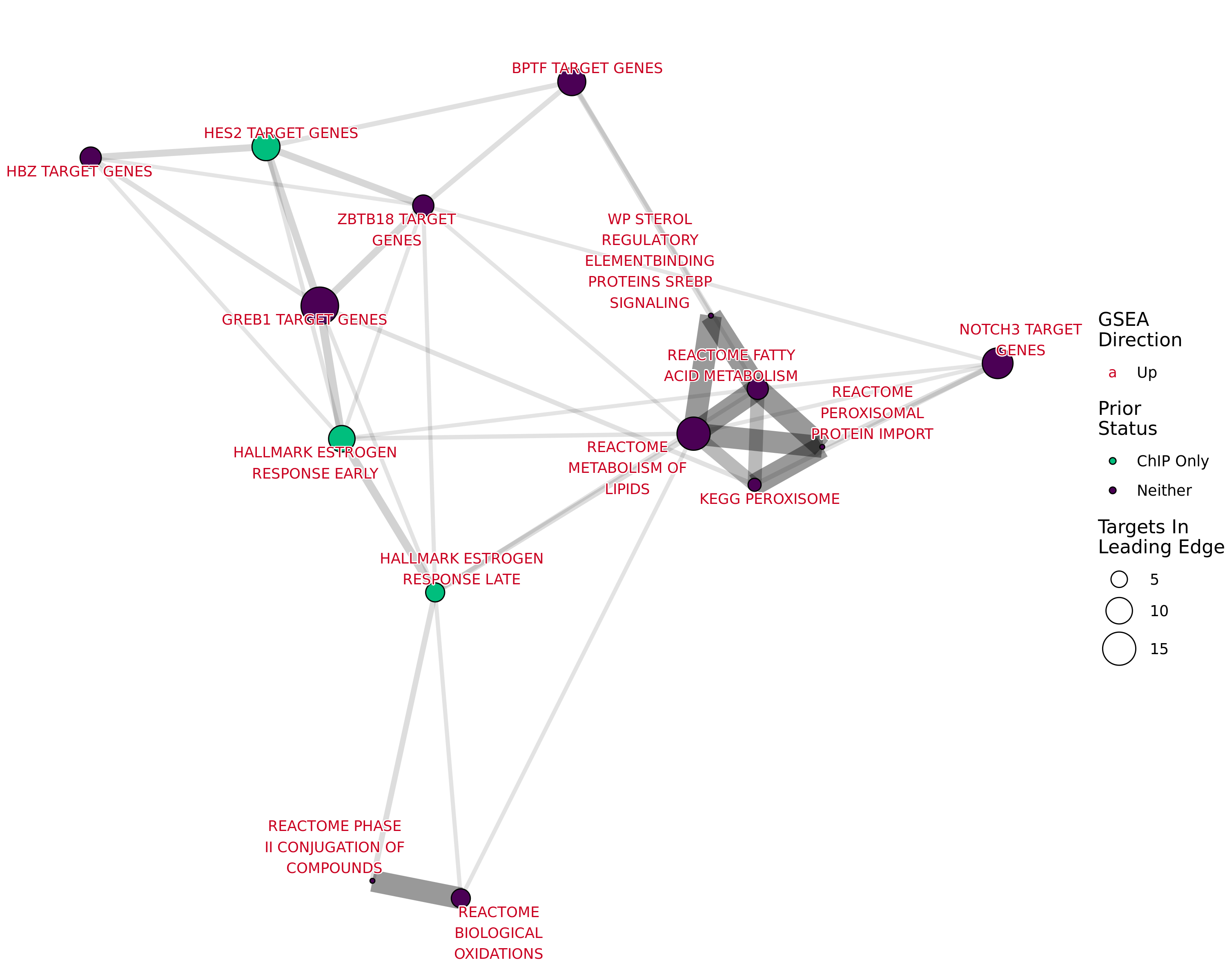
Pathway & Functional Analysis
- Look for biologically relevant signals in DE genes or network modules
- Enriched pathways
- Common transcription factors
- Drug target signals
- Compare to public datasets
(if appropriate)
- Interpretation is key
- Researchers love to invent stories
Acknowledgements
TKI / BODL Nerds
- Alex Brown
- Jimmy Breen
- Sam Buckberry
- Liza Kretzschmar
- Yassine Souilmi
- Bastien Llamas
- Holly Martin
- Claudia Floreani
- Natasha Howard
- Anelle du Preez
- Kaashifah Bruce
- Amanda Satour-Richards
- Justine Clarke
- Katharine Brown
NCIG
- Hardip Patel
ALIGN
- Johanna Barclay
- Annalee Stearne
- Louise Lyons
Students
- Nhi Hin
- Jacqueline Rehn
- Nora Liu
- Lachlan Baer
- Megan Monaghan
- Monica Guilhaus
Bioinformatics Hub
- David Adelson
- Gary Glonek
- Dan Kortschak
- Nathan Watson-Haigh
- Rick Tearle
- Hien To
Additional Material
The Central Dogma
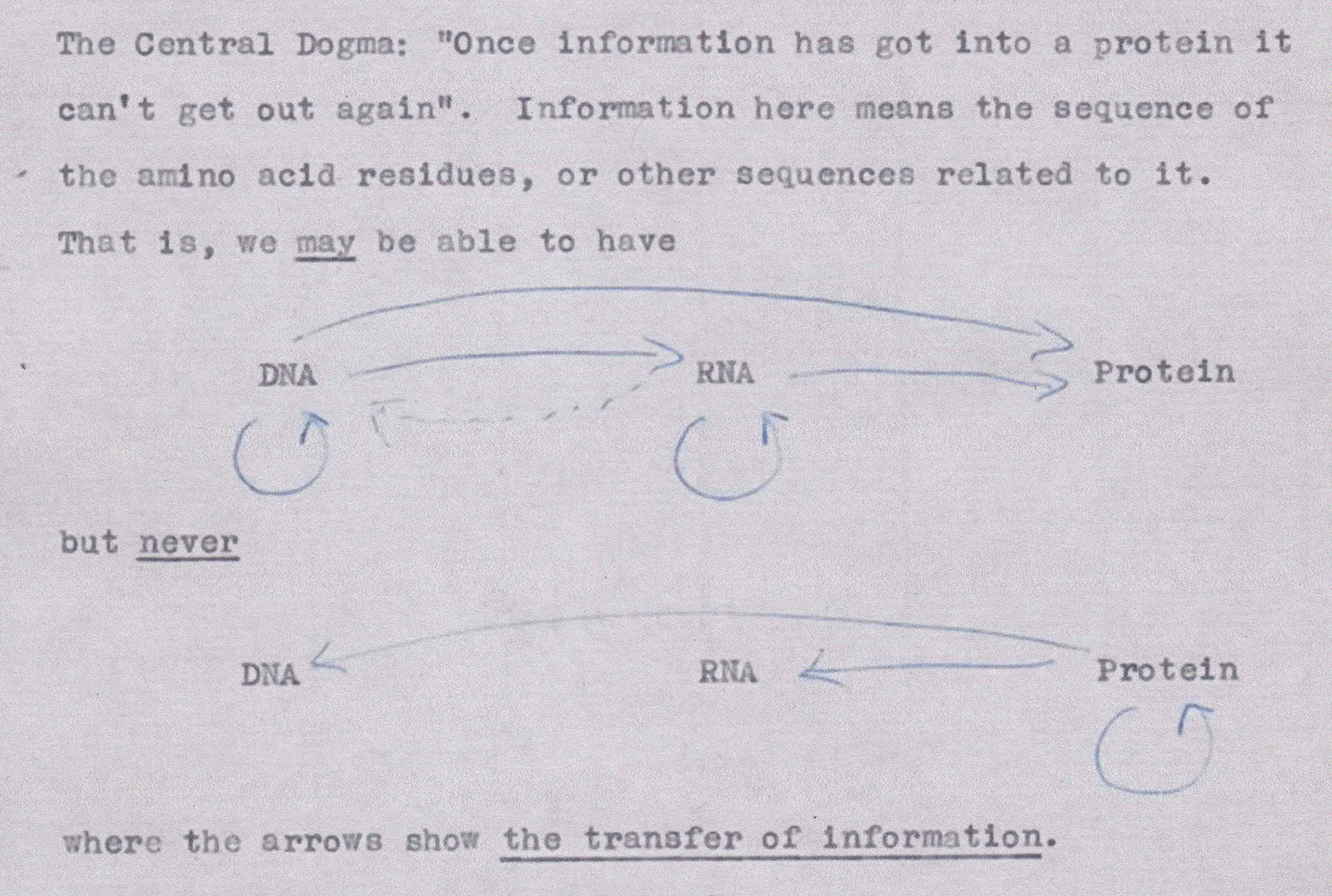
From Frances Crick, Ideas on Protein Synthesis, Unpublished Note, Wellcome Library, 1956
SEC14L2: A Random Example
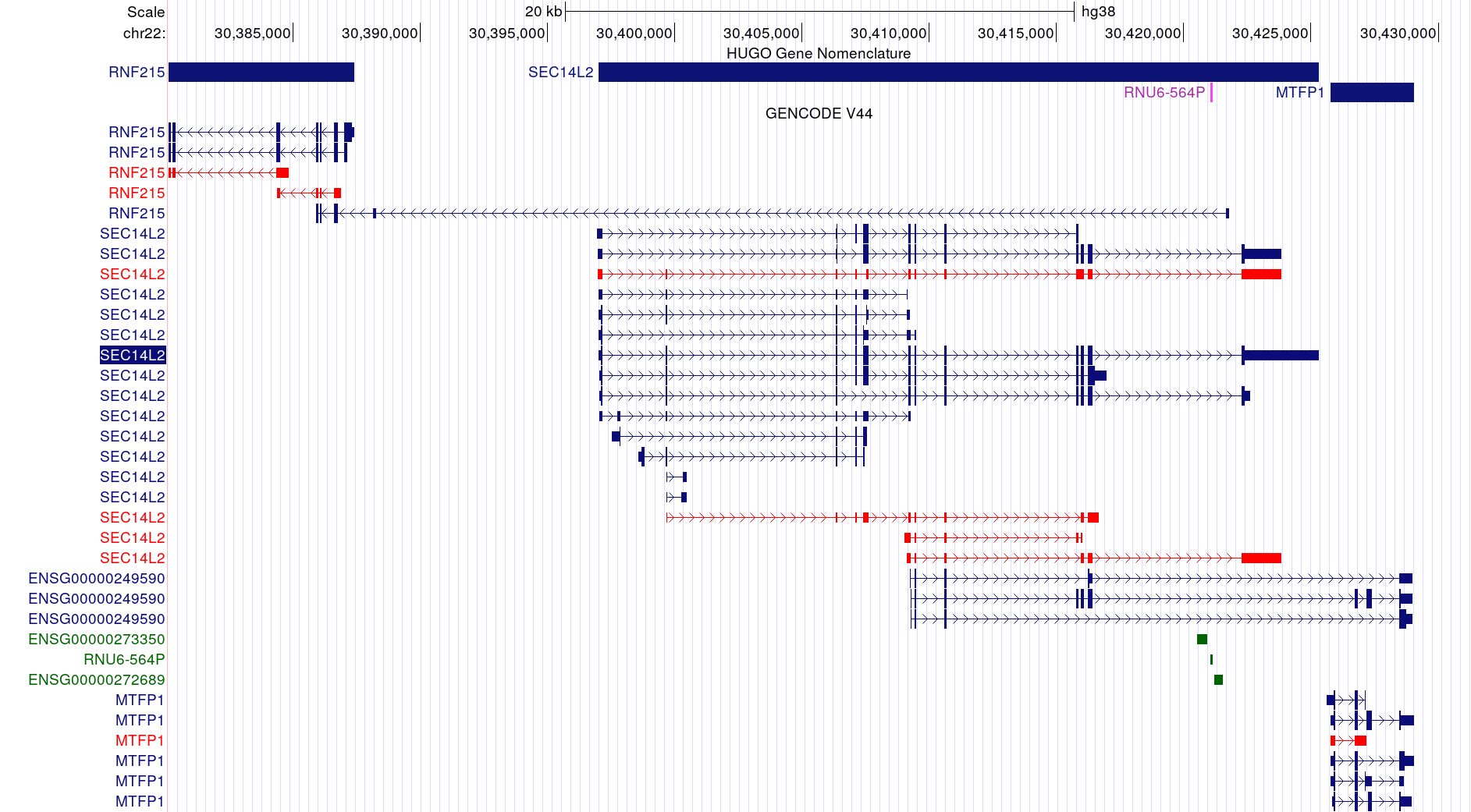
Long Reads
- Short read technology has ⇩⇩⇩ error rates
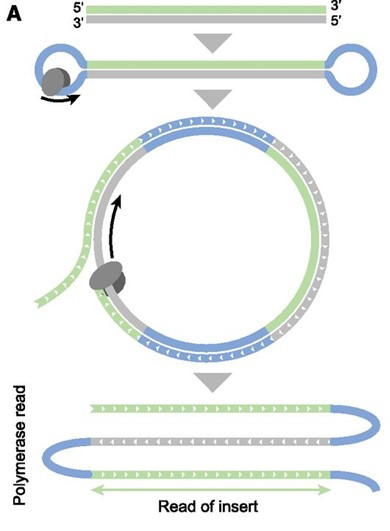
- Long reads have ⇧⇧⇧ error rates
- Reads are circularised
\(\implies\) errors corrected by repeat reads
- Great for identification of novel transcripts
\(\implies\) difficult to quantify- Creation of a custom reference transcriptome
- Challenging to refer back to functional annotations in reference

 shape during cell division
shape during cell division