extraChIPs: Going Beyond GRAVI
Black Ochre Data Laboratories, Telethon Kids Institute
March 17, 2023
Who Am I
Professionally
- 2022- Post-doctoral Bioinformatician: Black Ochre Data Laboratories
- 2020-2022 Dame Roma Mitchell Cancer Research Laboratories
- 2014-2020 Bioinformatics Hub, University of Adelaide
- 2008-2018 PhD Candidate, Barry Immunology Group
- 1991-2008 Musician
Research Interests
- Transcriptomics, Gene Regulation, Applied Statistics
- R programming, Bioconductor Nerd
Bioconductor Packages
ngsReports![]()
- Parse & plot output from
FastQC,cutadapt,STAR,macs2etc.
- Parse & plot output from
extraChIPs![]()
- ChIP-Seq Analysis & Visualisation
- Manipulation of
GRangesobjects - Built as infrastructure for the
GRAVIworkflow
But first…

extraChIPs
extraChIPs
- A walk-through of the package
- Usage well beyond ChIP-Seq data
- Approaches to detection of Differential ChIP-Seq Signal
extraChIPscurrently up on Bioconductor- Latest version is currently only on github
- Bioconductor 3.17 Release on 26th April
- R 4.3.0 on Friday 21st April
Utility Functions
Utility Functions
- Genomic Ranges objects (
GRanges) are analogous to bed files - Most functions for
GRangesfocus on the ranges component - My focus was on the data within the
mcols()element
- How can we play nicely with the
tidyverse?- Not treading on the toes of
plyranges - \(\implies\) enhancing approaches implemented by
plyranges
- Not treading on the toes of
tibble Coercion
## S3 method for class 'DataFrame'
as_tibble(x, rangeAsChar = TRUE, ...)
## S3 method for class 'GenomicRanges'
as_tibble(x, rangeAsChar = TRUE, name = "range", ...)
## S3 method for class 'Seqinfo'
as_tibble(x, ...)
## S3 method for class 'GInteractions'
as_tibble(x, rangeAsChar = TRUE, suffix = c(".x", ".y"), ...)- Very handy for interacting with
ggplot2anddplyretc - Handles
S4Compressed list columns well (so far)- Uses
vctrs::vec_proxy()to coerce to S3 lists
- Uses
- Reverse coercion also implemented
tibble Coercion
Starting with the protein-coding transcripts for CTLA4
GRanges object with 4 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <character>
[1] chr2 204732494-204738688 + | CTLA4 CTLA4-205
[2] chr2 204732666-204737498 + | CTLA4 CTLA4-201
[3] chr2 204732666-204737535 + | CTLA4 CTLA4-204
[4] chr2 204732666-204737535 + | CTLA4 CTLA4-203
-------
seqinfo: 24 sequences from GRCh37 genomeSeqinfo object with 24 sequences from GRCh37 genome:
seqnames seqlengths isCircular genome
chr1 249250621 FALSE GRCh37
chr2 243199373 FALSE GRCh37
chr3 198022430 FALSE GRCh37
chr4 191154276 FALSE GRCh37
chr5 180915260 FALSE GRCh37
... ... ... ...
chr20 63025520 FALSE GRCh37
chr21 48129895 FALSE GRCh37
chr22 51304566 FALSE GRCh37
chrX 155270560 FALSE GRCh37
chrY 59373566 FALSE GRCh37tibble Coercion
Now perform the coercion using as_tibble()
Coerce back to a GRanges uses colToRanges()
GRanges object with 4 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <character>
[1] chr2 204732494-204738688 + | CTLA4 CTLA4-205
[2] chr2 204732666-204737498 + | CTLA4 CTLA4-201
[3] chr2 204732666-204737535 + | CTLA4 CTLA4-204
[4] chr2 204732666-204737535 + | CTLA4 CTLA4-203
-------
seqinfo: 24 sequences from GRCh37 genome*MC() Functions
- Many
GRangesfunctions just operate on the rangereduce,intersect,setdiff,union
extraChIPsvariants also return themcolselementreduceMC,intersectMC,setdiffMC,unionMCdistinctMCandchopMCreplicatetidyversefunctions
- There is a performance overhead… 😞
reduceMC()
Say we wish to find the TSS for CTLA transcripts
GRanges object with 4 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <character>
[1] chr2 204732494 + | CTLA4 CTLA4-205
[2] chr2 204732666 + | CTLA4 CTLA4-201
[3] chr2 204732666 + | CTLA4 CTLA4-204
[4] chr2 204732666 + | CTLA4 CTLA4-203
-------
seqinfo: 24 sequences from GRCh37 genomereduceMC()
Say we wish to find the TSS for CTLA transcripts
GRanges object with 4 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <character>
[1] chr2 204732494 + | CTLA4 CTLA4-205
[2] chr2 204732666 + | CTLA4 CTLA4-201
[3] chr2 204732666 + | CTLA4 CTLA4-204
[4] chr2 204732666 + | CTLA4 CTLA4-203
-------
seqinfo: 24 sequences from GRCh37 genome
reduceMC() returns the mcols as well
GRanges object with 2 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <CharacterList>
[1] chr2 204732494 + | CTLA4 CTLA4-205
[2] chr2 204732666 + | CTLA4 CTLA4-201,CTLA4-204,CTLA4-203
-------
seqinfo: 24 sequences from GRCh37 genomeintersectMC()
- Find the intersecting range for a ChIP-Seq peak
GRanges object with 1 range and 0 metadata columns:
seqnames ranges strand
<Rle> <IRanges> <Rle>
[1] chr2 204732401-204732600 *
-------
seqinfo: 24 sequences from GRCh37 genomeGRanges object with 1 range and 0 metadata columns:
seqnames ranges strand
<Rle> <IRanges> <Rle>
[1] chr2 204732494-204732600 *
-------
seqinfo: 24 sequences from GRCh37 genome- NB:
subsetByOverlaps()would return the entire initial range - Only the
mcolsfrom the query range are returned
setdiffMC()
- Here we can chop out ranges
- Find the region of CTLA4 not covered by the 3 shorter transcripts
GRanges object with 4 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <character>
[1] chr2 204732494-204738688 + | CTLA4 CTLA4-205
[2] chr2 204732666-204737498 + | CTLA4 CTLA4-201
[3] chr2 204732666-204737535 + | CTLA4 CTLA4-204
[4] chr2 204732666-204737535 + | CTLA4 CTLA4-203
-------
seqinfo: 24 sequences from GRCh37 genomeGRanges object with 2 ranges and 2 metadata columns:
seqnames ranges strand | gene_name transcript_name
<Rle> <IRanges> <Rle> | <character> <character>
[1] chr2 204732494-204732665 + | CTLA4 CTLA4-205
[2] chr2 204737536-204738688 + | CTLA4 CTLA4-205
-------
seqinfo: 24 sequences from GRCh37 genome- Again, we retain the
mcolsinformation
distinctMC()
- Very handy after a
plyranges::join_*()with multiple matches
GRanges object with 2 ranges and 1 metadata column:
seqnames ranges strand | prop_methylated
<Rle> <IRanges> <Rle> | <numeric>
[1] chr2 204732499 * | 0.20
[2] chr2 204732562 * | 0.99
-------
seqinfo: 24 sequences from GRCh37 genome- The join will expand to both methylated sites
distinctMC()
- Very handy after a
plyranges::join_*()with multiple matches
GRanges object with 2 ranges and 1 metadata column:
seqnames ranges strand | prop_methylated
<Rle> <IRanges> <Rle> | <numeric>
[1] chr2 204732499 * | 0.20
[2] chr2 204732562 * | 0.99
-------
seqinfo: 24 sequences from GRCh37 genome- We can just keep the first mapping after sorting by some parameter
peak %>%
plyranges::join_overlap_left(meth) %>%
arrange(1/(prop_methylated)) %>%
distinctMC(.keep_all = TRUE)GRanges object with 1 range and 1 metadata column:
seqnames ranges strand | prop_methylated
<Rle> <IRanges> <Rle> | <numeric>
[1] chr2 204732401-204732600 * | 0.99
-------
seqinfo: 24 sequences from GRCh37 genomechopMC()
- Wraps
tidyr::chop()- Chops by any specified column, always including the range
- Could then run
vapply()on the column to summarise
GRanges object with 1 range and 1 metadata column:
seqnames ranges strand | prop_methylated
<Rle> <IRanges> <Rle> | <NumericList>
[1] chr2 204732401-204732600 * | 0.20,0.99
-------
seqinfo: 1 sequence from an unspecified genome; no seqlengths
Visualisation
Visualisations
extraChIPswas written for theGRAVIworkflow- Added functions for common visualisations in the context of ChIP-Seq
- Commonly use a
SummarizedExperimentobject with multiple ‘assays’- Usually have a
rowRangeselement
- Usually have a
- Most plotting functions built for
GRanges/data.frame-like objects
Visualisations
plotAssay*()for inspection of pre-analysed dataplotAssayDensities(),plotAssayPCA(),plotAssayRle()
plotOverlaps(),plotPie()andplotSplitDonut()- Compare sets of ranges using
Venn/UpSet, Pie or Donut charts
- Compare sets of ranges using
plotProfileHeatmap()- Visualise binding intensities at multiple sites/samples
plotHFGC(): plot HiC, Features, Genes, Coverage- All elements are optional
- Define your objects \(\implies\) keep running across multiple ranges
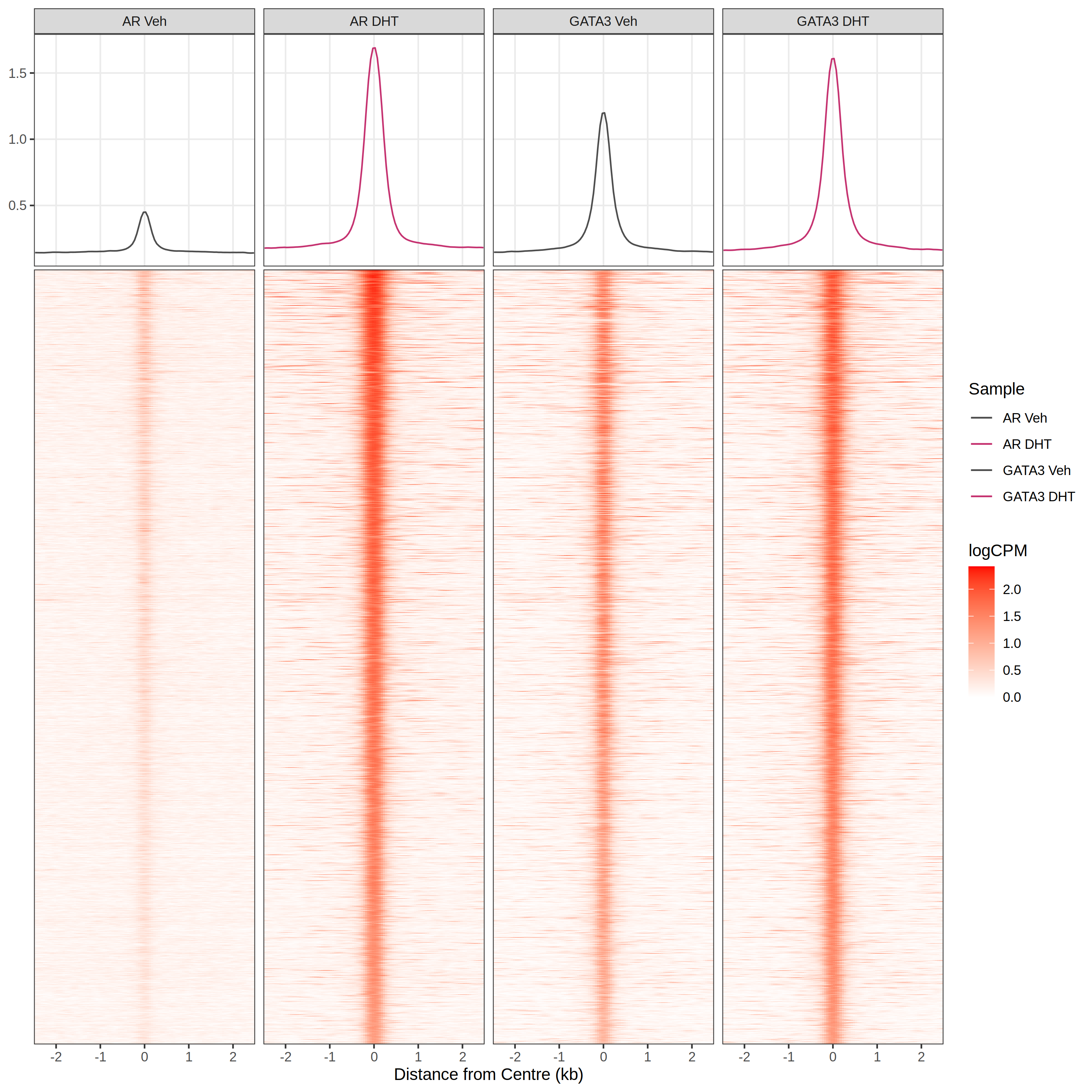
plotProfileHeatmap()
- Provide a set of ranges
- Centre & standardise width
- Read in data from
BigWigFileListgetProfileData(bwfl, gr)
- Plot data
- Uses
ggplot2for easy customisation- Top panel plots using
ggside
- Top panel plots using
- Still frustratingly slow… 😞
plotHFCG()
- Wraps plotting functions from
Gviz - HiC as
GenomicInteractionsobject - Each Feature track as a
GRangeswithin a named list - Each Gene Track as a
GRangeswithin a named list - Each coverage track is a BigWigFileList
- Pass a list of
BigWigFileListobjects for multiple tracks - Coverage tracks can also have
GRangesto annotate them
- Pass a list of
plotHFGC()
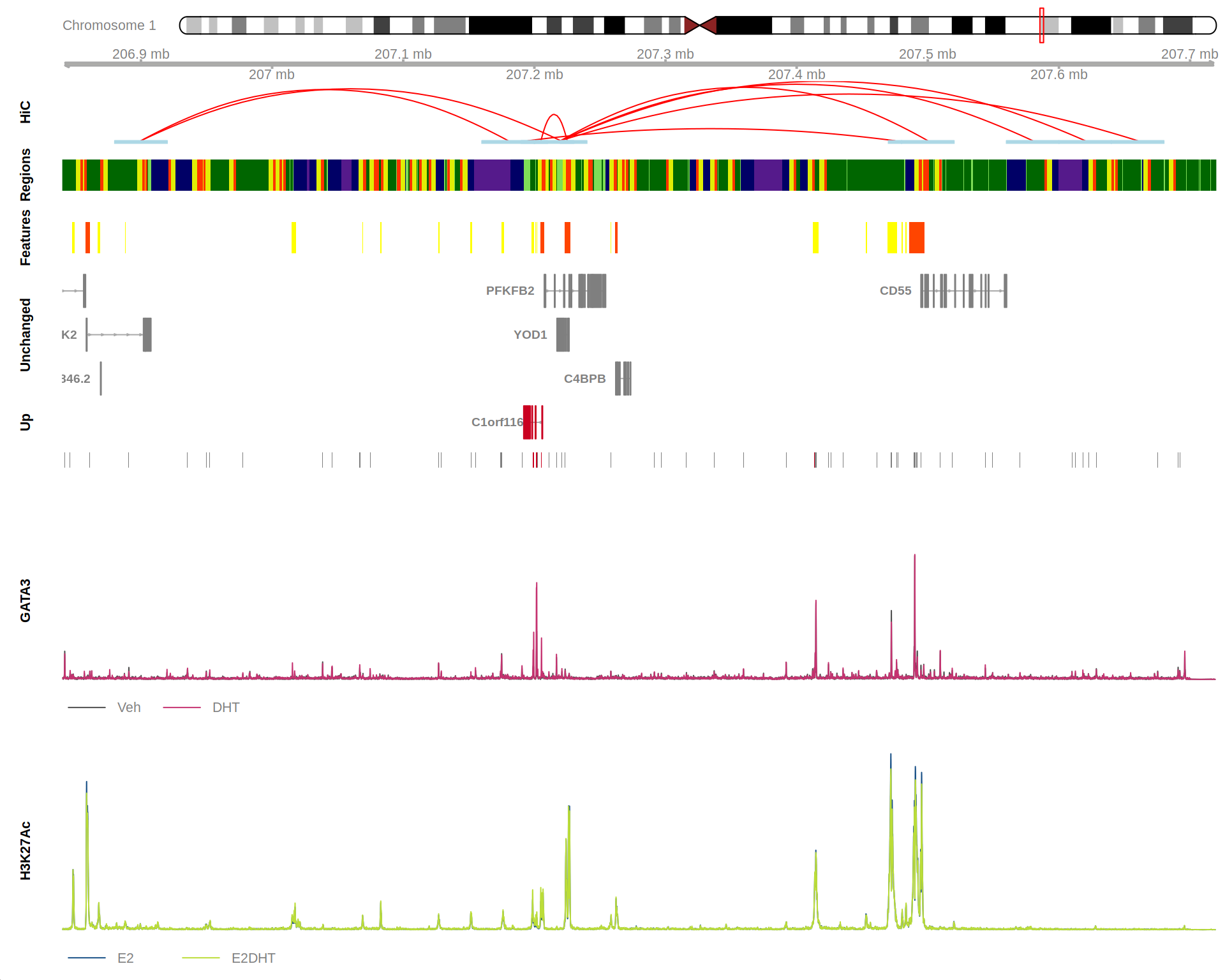
Here I’m checking binding for a DE gene (C1orf116)…
plotPie()
plotPie(consensus_peaks, fill = "region") +
scale_fill_manual(values = colours$regions) +
theme(legend.position = "none")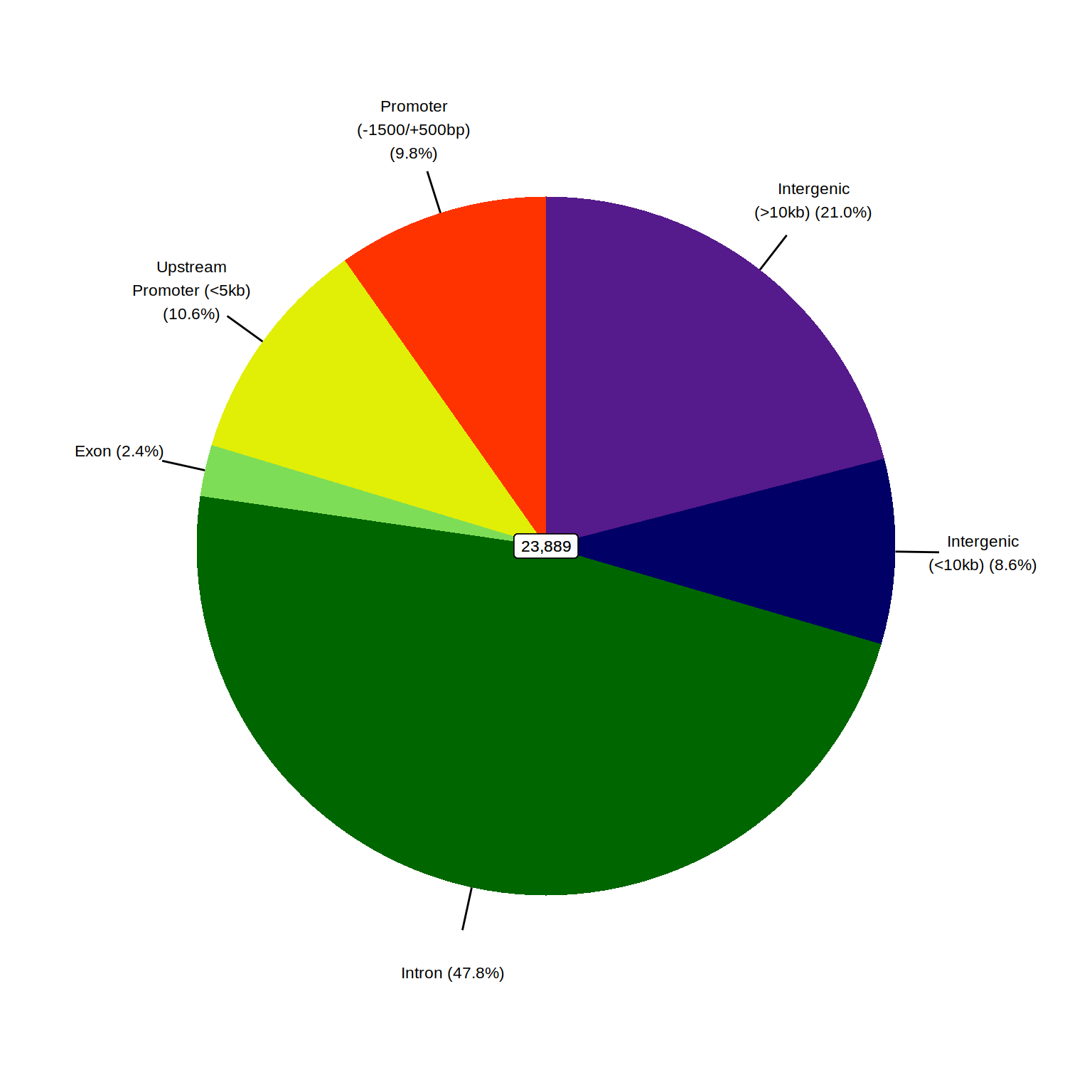
AR binding sites mapped to genomic regions
plotSplitDonut()
- Compares two columns from a
GRangesordata.frameobject - Can set inner & outer palettes separately
- Huge flexibility in labelling
- Uses
patchworkto drive the layout - Getting legend positions right is still a bit tricky (i.e. hacky…)
plotSplitDonut()
plotSplitDonut(
object, inner, outer, scale_by = NULL,
r_centre = 0.5, r_inner = 1, r_outer = 1,
total_size = 5, total_glue = "{comma(N)}",
inner_glue = "{inner} {.data[[inner]]}\n{percent(p,0.1)}",
outer_glue = "{outer} {.data[[outer]]}\n{percent(p,0.1)}",
inner_label = c("label", "text", "none"),
outer_label = c("label", "text", "none"),
label_alpha = 1, inner_label_alpha = NULL, outer_label_alpha = NULL,
label_size = 3, inner_label_size = NULL, outer_label_size = NULL,
min_p = 0.05, inner_min_p = NULL, outer_min_p = NULL,
explode_inner = NULL, explode_outer = NULL, explode_query = c("AND", "OR"),
explode_x = 0, explode_y = 0, explode_r = 0, nudge_r = 0.5,
expand = 0.1,
inner_palette = NULL, outer_palette = NULL,
inner_legend = TRUE, outer_legend = TRUE,
layout = c(main = area(1, 1, 6, 6), lg1 = area(2, 7), lg2 = area(4, 7)), ...
)- Need to add
label& tweak how label size is set globally
plotSplitDonut()
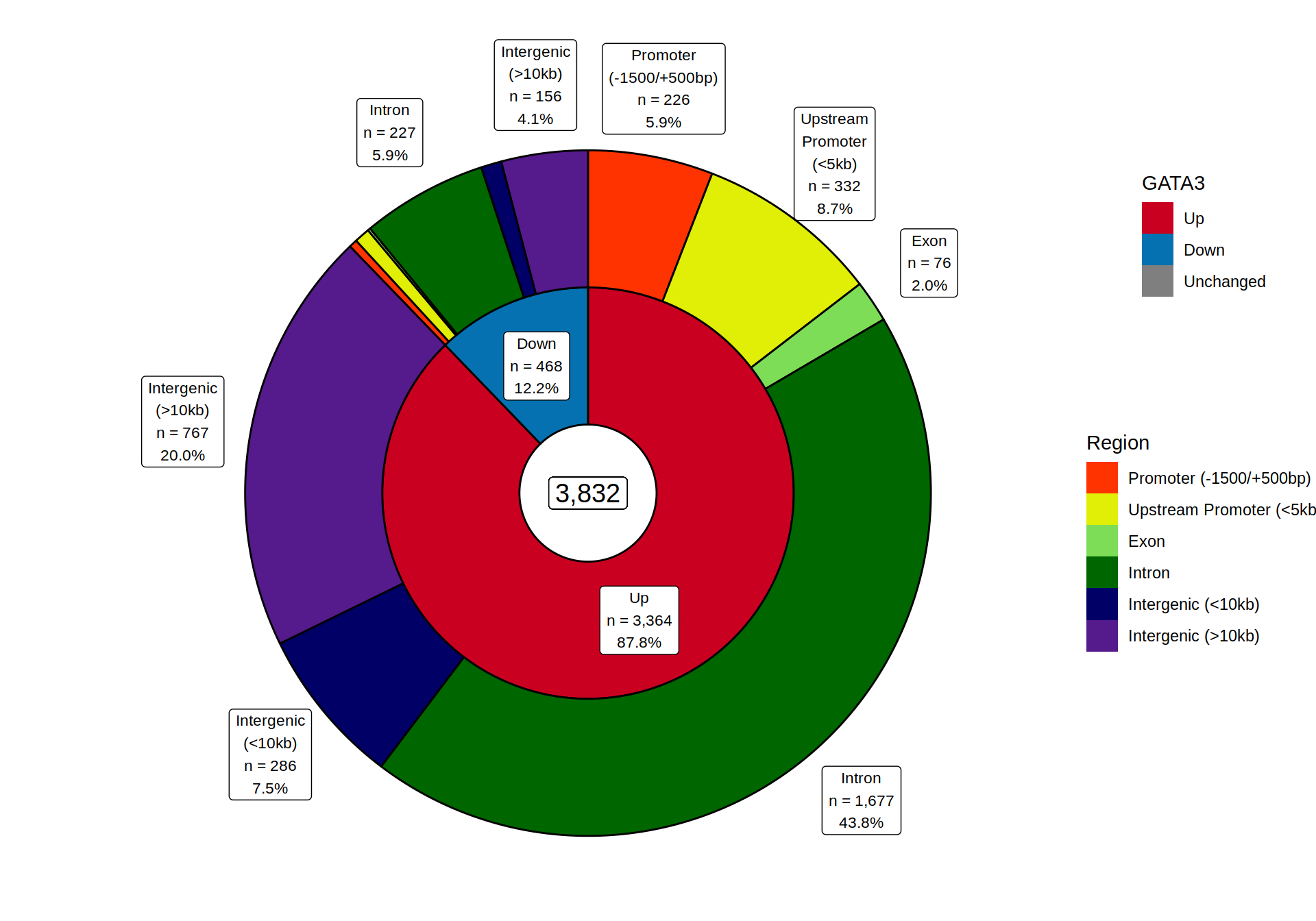
target <- "GATA3
merged_results %>%
plotSplitDonut(
inner = target, outer = "Region", min_p = 0.01,
inner_glue = "{.data[[inner]]}\nn = {comma(n, 1)}\n{percent(p, 0.1)}",
outer_glue = "{str_wrap(.data[[outer]], 12)}\nn = {comma(n, 1)}\n{percent(p, 0.1)}",
inner_palette = direction_colours, outer_palette = region_colours
)
plotSplitDonut()
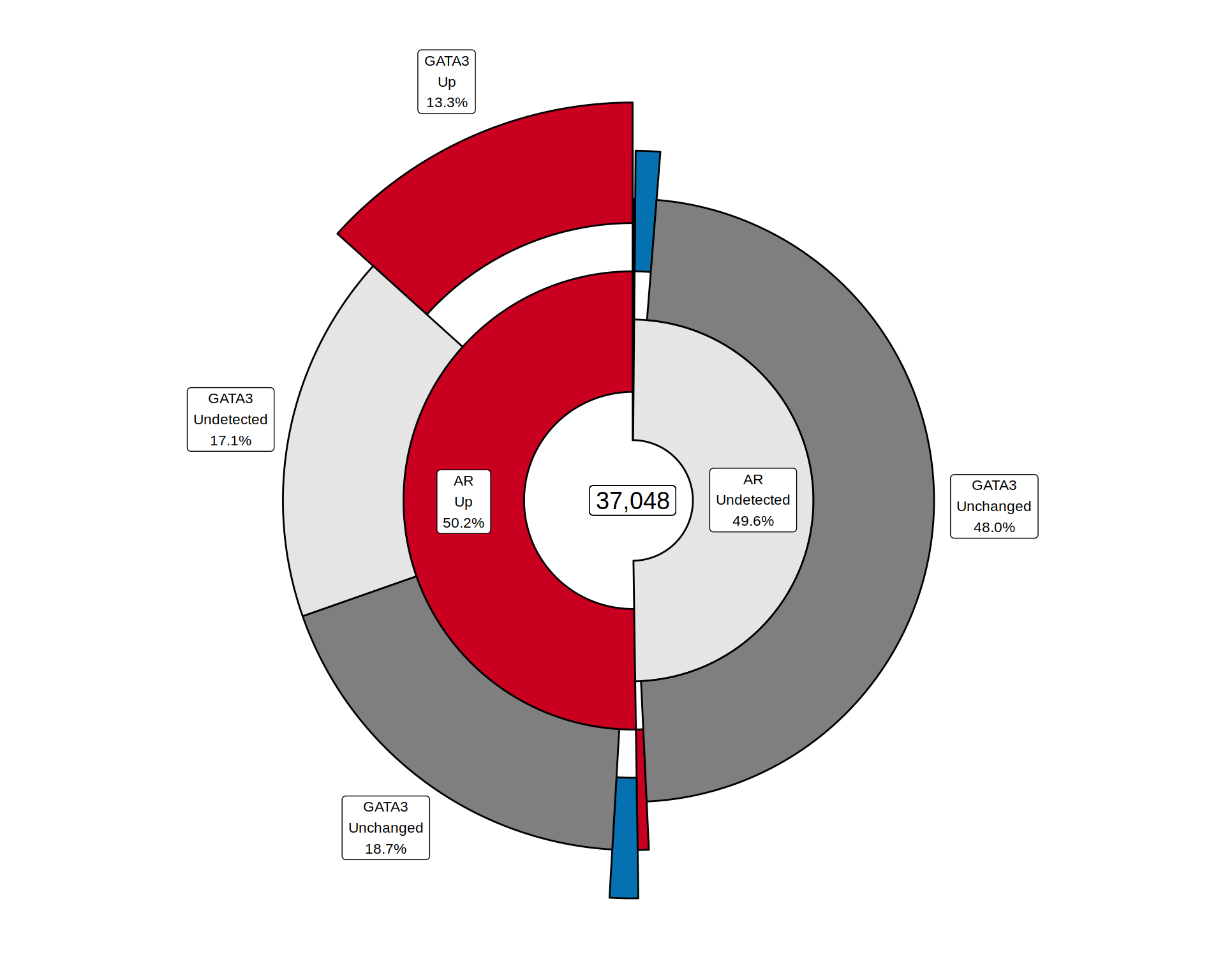
all_windows %>%
plotSplitDonut(
inner = colnames(.)[[1]], outer = colnames(.)[[2]], min_p =0.02,
inner_glue = "{inner}\n{.data[[inner]]}\n{percent(p, 0.1)}",
outer_glue = "{outer}\n{.data[[outer]]}\n{percent(p, 0.1)}",
explode_inner = "Down|Up", explode_outer = "Down|Up", explode_r = 0.4,
explode_query = "OR", nudge_r = 0.5,
inner_palette = colours$direction, outer_palette = colours$direction
inner_legend = FALSE, outer_legend = FALSE
) 
plotOverlaps()
- Calls 1)
ComplexUpset::upset()or 2)VennDiagram::draw.*wise.venn() - Size-scaled Venn Diagrams are tricky in
R…
ex <- list(x = letters[1:5], y = letters[c(6:15, 26)], z = letters[c(2, 10:25)])
plotOverlaps(ex, type = "upset", set_col = 1:3, labeller = stringr::str_to_title)
Differential ChIP Signal
ChIP-Seq
- A protein of interest is:
- Cross-linked to DNA
- Immuno-Precipitated using antibody for ChIP target
- Cross-links reversed \(\implies\) DNA sequenced
- Sequences should be enriched for DNA-bound regions
- ChIP targets can be transcription factors, histone marks (e.g. H3K27ac)
- TFs are generally shorter (300-500bp) regions
- Histone marks can be far broader (>1kb)
- Can be quantitative or have informative sequences
Differential Signal/Binding
- Looking for increase/decrease in binding (or signal) at a genomic region
- Fundamentally different to RNA-Seq:
- Only 0/1/2 fragments per cell
- Increase in signal \(\implies\) Target bound in more cells at that site
Differential Binding
- Forms a key part of the
GRAVIworkflow GRAVIis to run multiple ChIP targets- Need a method which is robust when automatically deployed
- Normalisation becomes important
- Can handle targets which are:
- Cytoplasmic \(\rightarrow\) Nuclear
- Partially sequestered \(\rightarrow\) More able to bind
- Just relocated on the DNA
- Also robust for H3K27ac marks etc
DRMCRL Data
Usually DHT Vs Veh or E2+DHT Vs E2
- AR shifts to nucleus with DHT-treatment
- AR relocates ER on the chromosome (Hickey et al. 2021)
- Does AR bring some/any GATA3 to the nucleus with it?
- How does this relate to H3K27ac signal changes
- Much broader peaks to transcription factors
- Include Differential H3K27ac signal alongside AR/ER/GATA3
Can I develop a differential signal workflow for all situations?
DiffBind
- Is the most common method for ChIP-Seq analysis (Ross-Innes et al. 2012)
- Uses defined peaks (e.g. macs2 consensus peaks)
- Re-centres peaks when loading
- Fixed width across all peaks (e.g. all 501bp)
- Defaults to:
- Library Size Normalisation, i.e
log(counts/lib.size) ~ X DESeq2(just like RNA-Seq)- Can also use
edgeR& choose alternative normalisation
- Library Size Normalisation, i.e
csaw
- Developed by Gordon Smyth & Aaron Lun (Lun and Smyth 2016)
- Uses sliding windows
- Avoids bias from counting within defined peaks (Lun and Smyth 2014)
- Regions defined independently of signal
- After counting across genome \(\implies\) throw away low/no signal windows
- No normalisation method by default (we choose)
- Statistical Test on remaining windows (
edgeR::glmQLF()) - Summarise tests across overlapping windows \(\implies\) merged region
- Tests/p-values are clearly dependent
- Can end up with tighter and broader regions than DiffBind
An Example
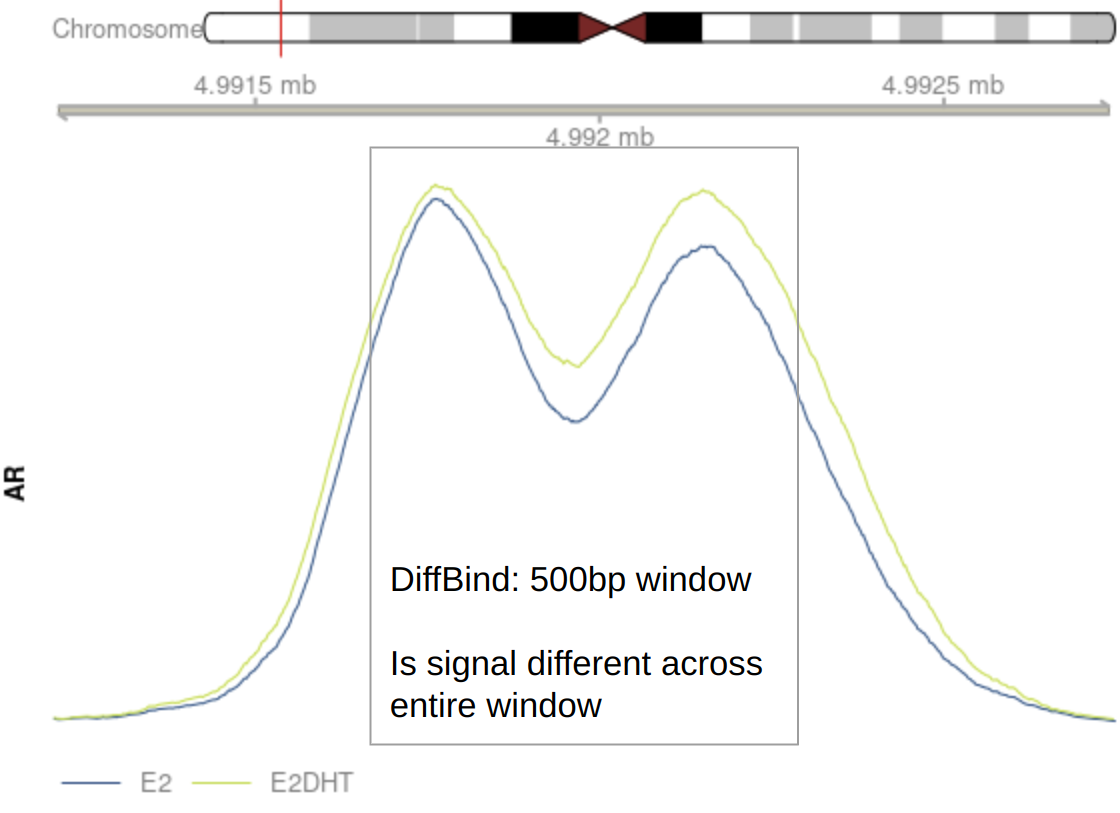
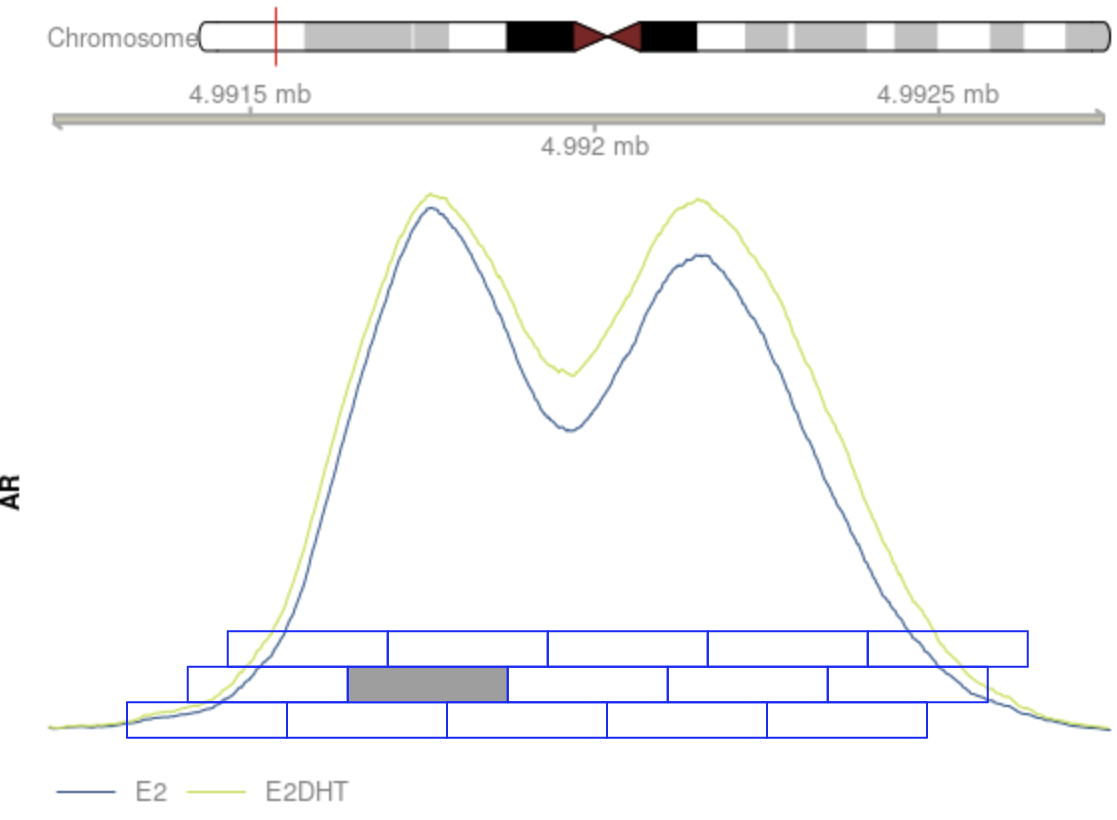
Use the maximal signal window?
Merging Windows
csaw offers multiple options
- Take window with highest signal as representative of entire region
- Choose the window with the most extreme logFC
- Often low signal windows
- Choose lowest p-value
- Needs careful adjustment after merging
- Use Simes’ Method to merge all \(p\)-values
- Chooses logFC from lowest p-value as representative
extraChIPs: Window Merging
- All
csawmethods wrapped for slightly different output structure
- Merge using harmonic-mean p-value (Wilson 2019)
- \(\text{hmp} =\frac{n}{\sum_i^np_i^{-1}}\) if no weights provided
- \(\text{hmp} =\frac{\sum_{i=1}^nw_i}{\sum_i^nw_i p_i^{-1}}\) if weights provided
- Actually uses the asymptotically exact harmonic mean
- Spent the weekend getting a 100x speed up from
harmonicmeanp
extraChIPs: Window Merging
- Robust to dependence
- Controls FDR within a set of tests
- Can also produce strict FWER controlled version for complete dataset
- Return weighted-mean of logFC/logCPM using \(p_i^{-1}\) for weights
- e.g. \(\widehat{\text{logFC}} = \frac{\sum_i^np_i^{-1}\text{logFC}_i}{\sum_i^np_1^{-1}}\)
extraChIPs: Window Merging
- Outperforms Simes’ consistently
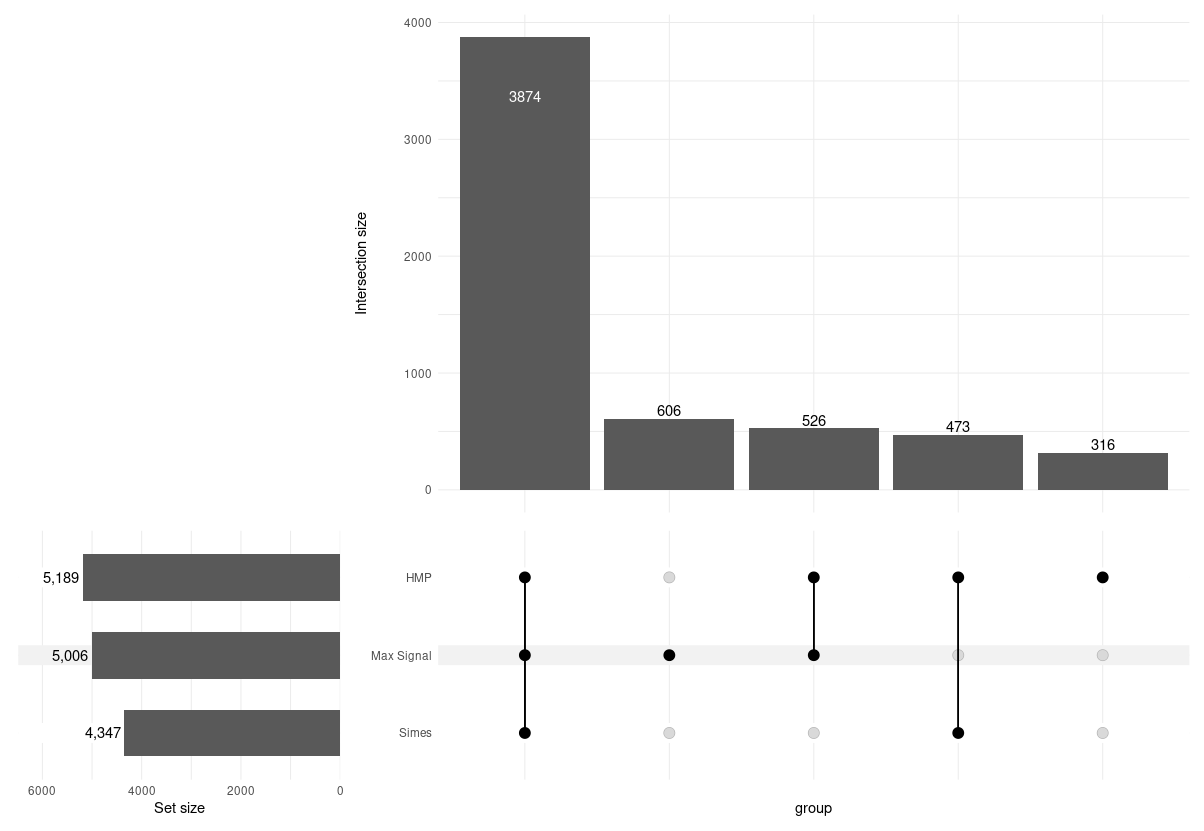
Results Example 1
A ~3kb H3K27ac region unique to harmonic mean
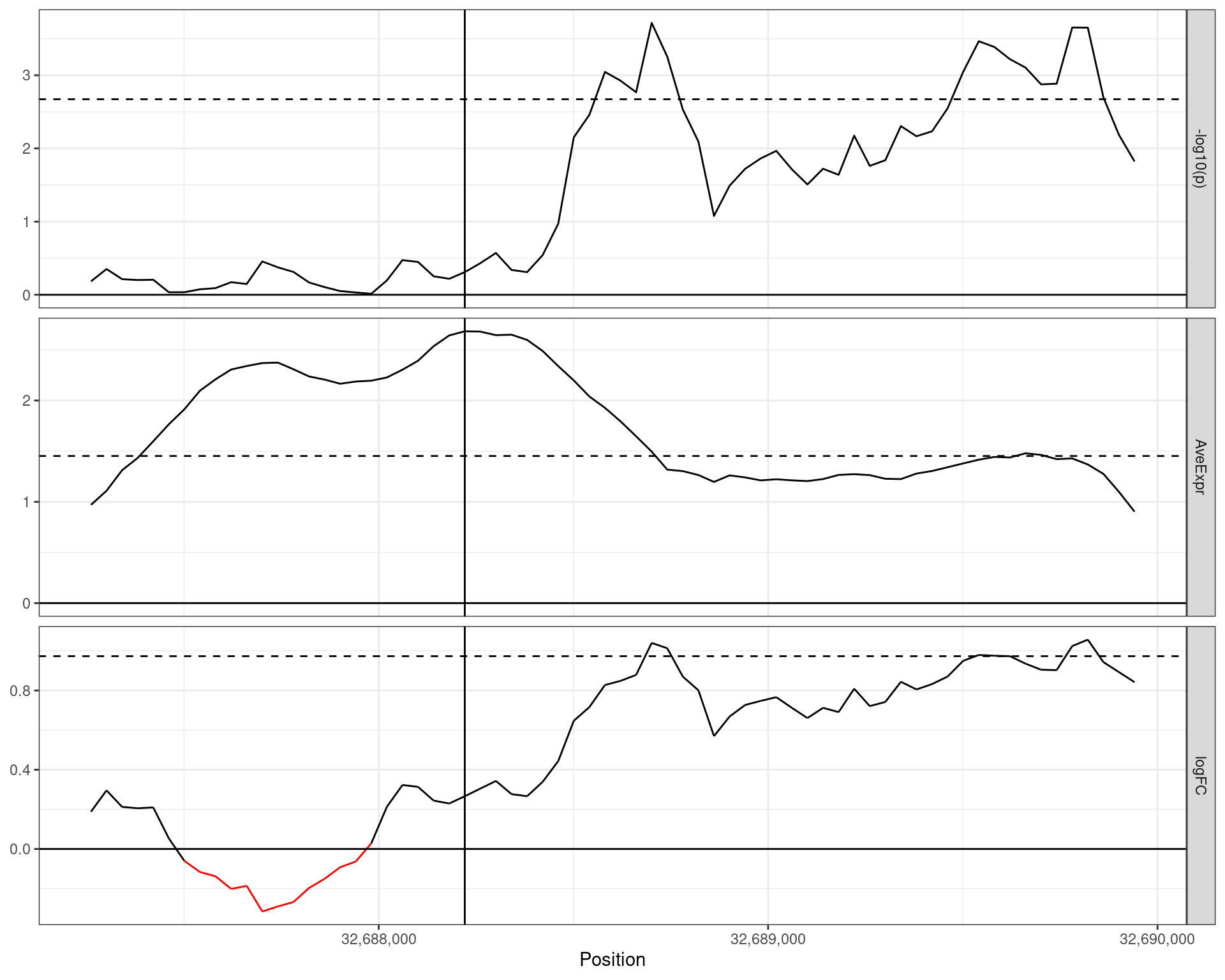
- \(-\log_{10}p\) in the top panel
- Dashed line is \(-\log_{10}(\text{hmp})\)
- Dashed line is \(-\log_{10}(\text{hmp})\)
- Average Signal is the middle panel
- Dashed line is weighted mean
- Dashed line is weighted mean
- logFC in bottom panel
- Dashed line is weighted mean
- Vertical line is maximal signal
Normalisation Methods
- Library Size only lacks significant power
- No attempt to handle technical variability
- TMM/RLE are both ok if overall signal is the same in all treatments
- DRMCRL data has pooled input:
normalise to input
- Is there a general method for automating in a workflow ?
My SuggestionL SQLT
- Calculate
logCPM- Effectively normalises by library size
- Smooth Quantile Normalisation (Stephanie C. Hicks et al. 2018)
- Generalisation of quantile normalisation within groups
- Reduces technical variability
limma-trend(Law et al. 2014) using normalised logCPM
- Merge Windows using Harmonic Mean \(p\)-value (
extraChIPs)
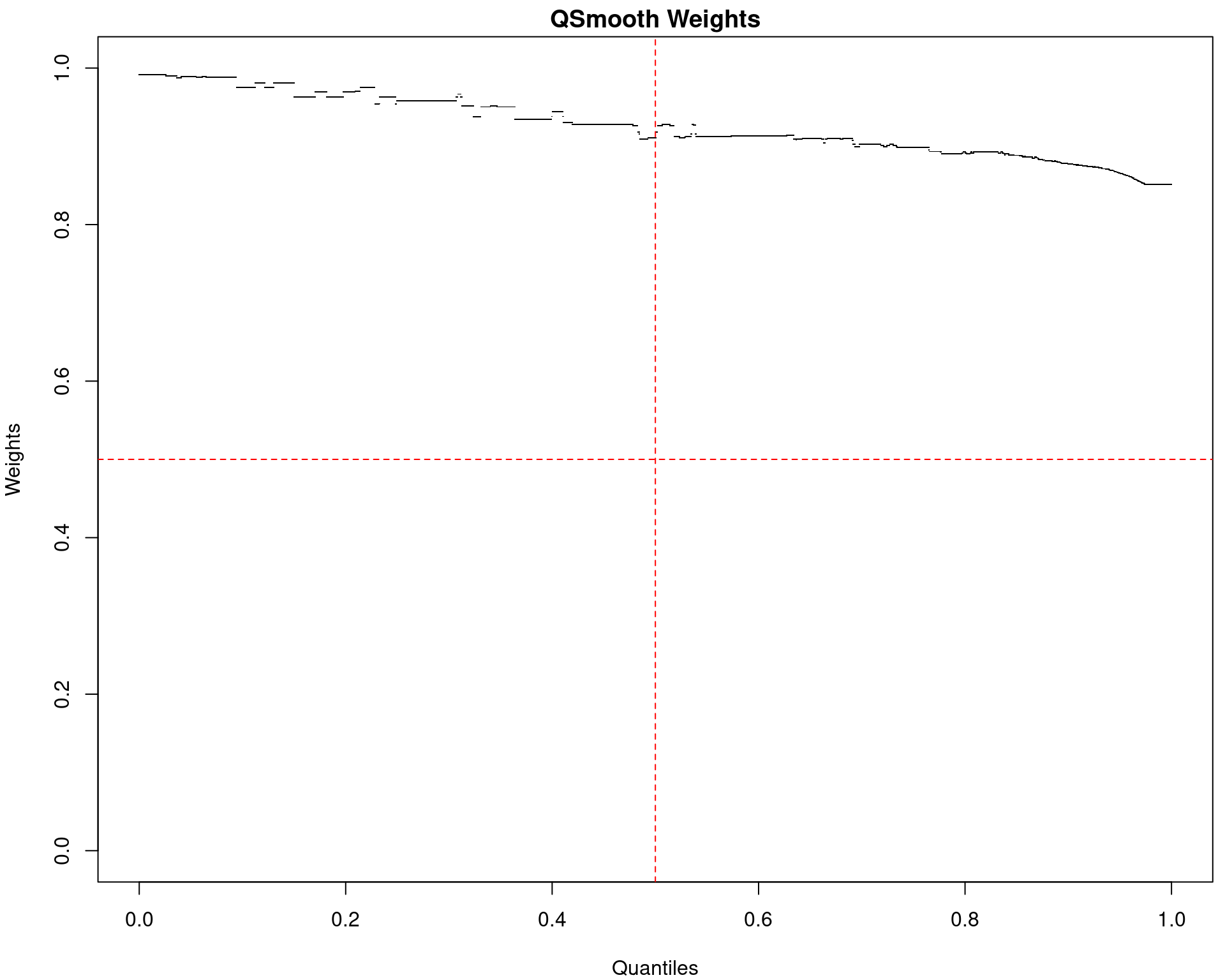
Smooth Quantile Normalisation
- Quantile Normalisation was standard practice for microarrays
- Gives all samples the identical distribution
- Reduces technical variability between samples
- Smooth-QN allows for treatment groups
- Uses weights to determine areas which diverge between groups
- Fits a weighted average of expected values
quantro(Stephanie C. Hicks and Irizarry 2015) provides a test for- Difference in distribution medians (\(F\)-test)
- Difference in overall distributions (permuted)
SQN
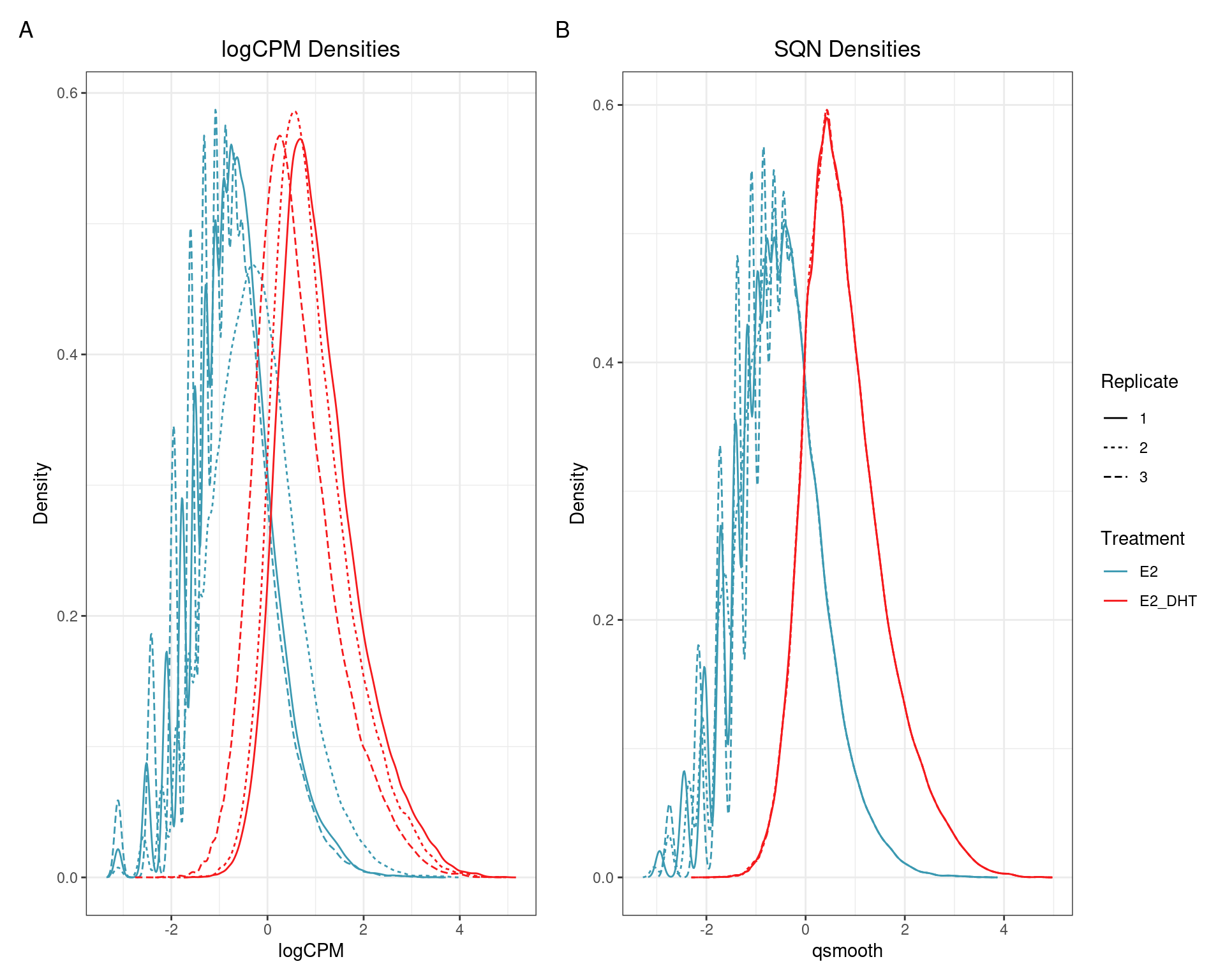
(quantro: \(p_{med} = 0.002\); \(p_{dist} < 5e4\))
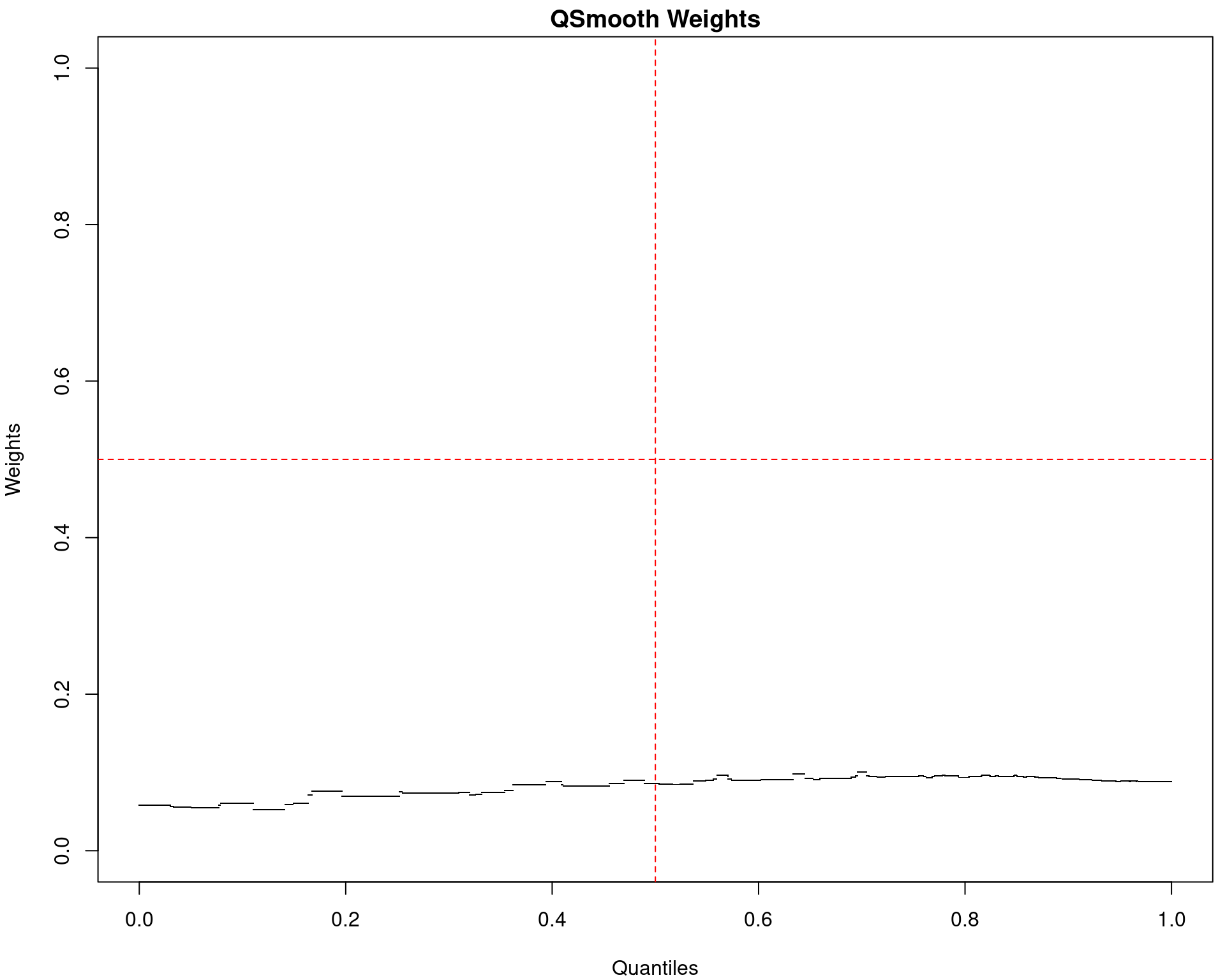
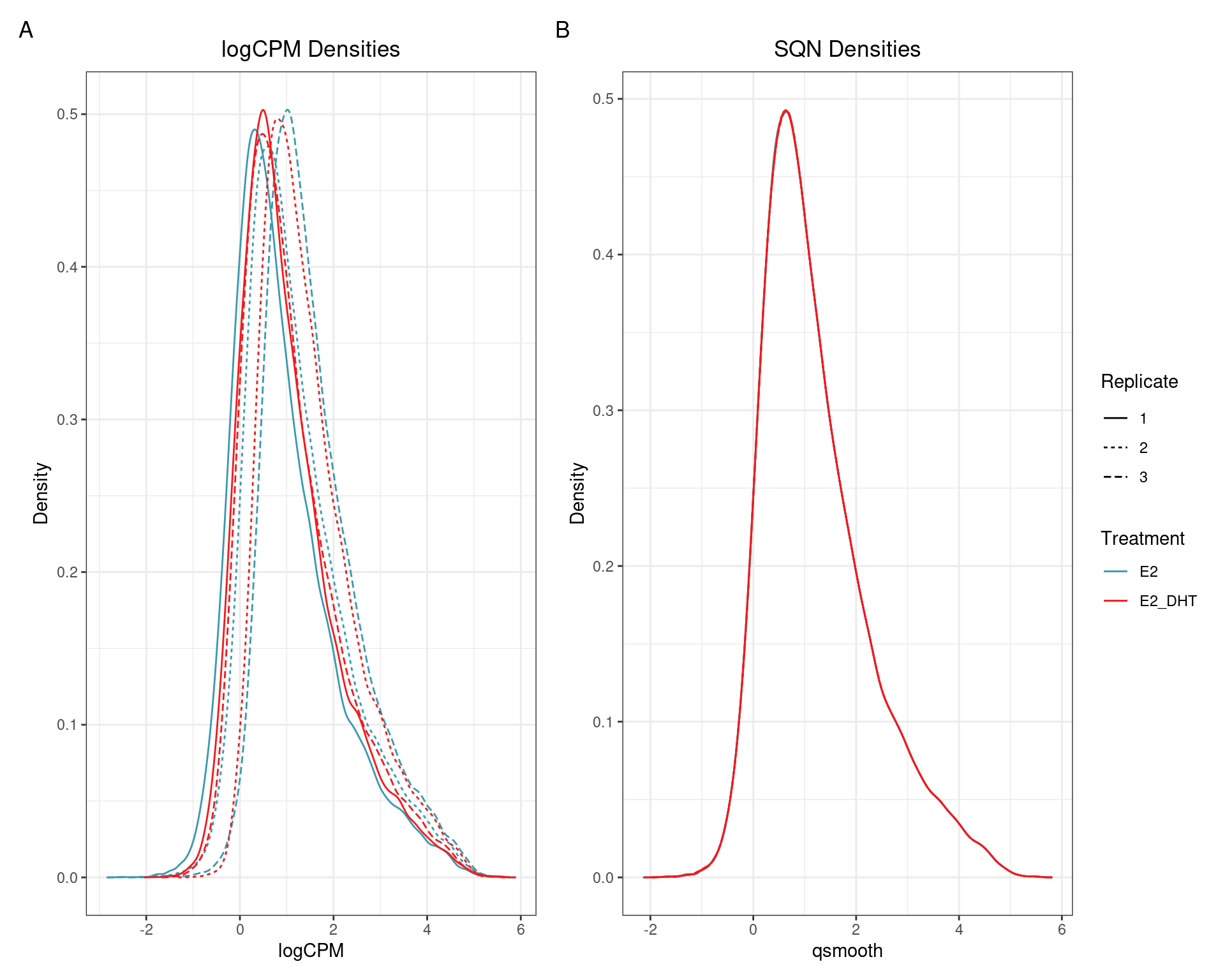
(quantro: \(p_{med} = 0.91\); \(p_{dist} = 0.42\))
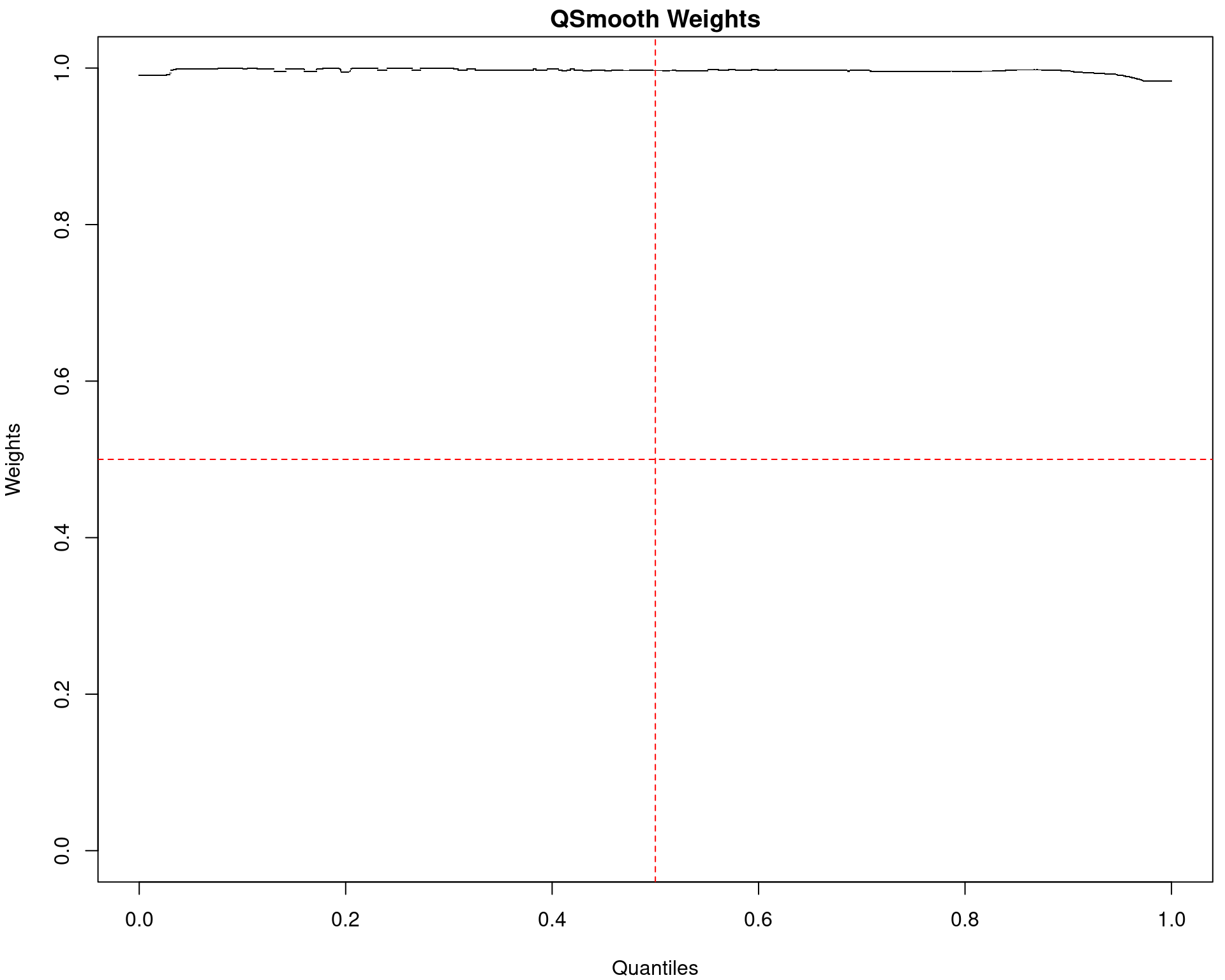
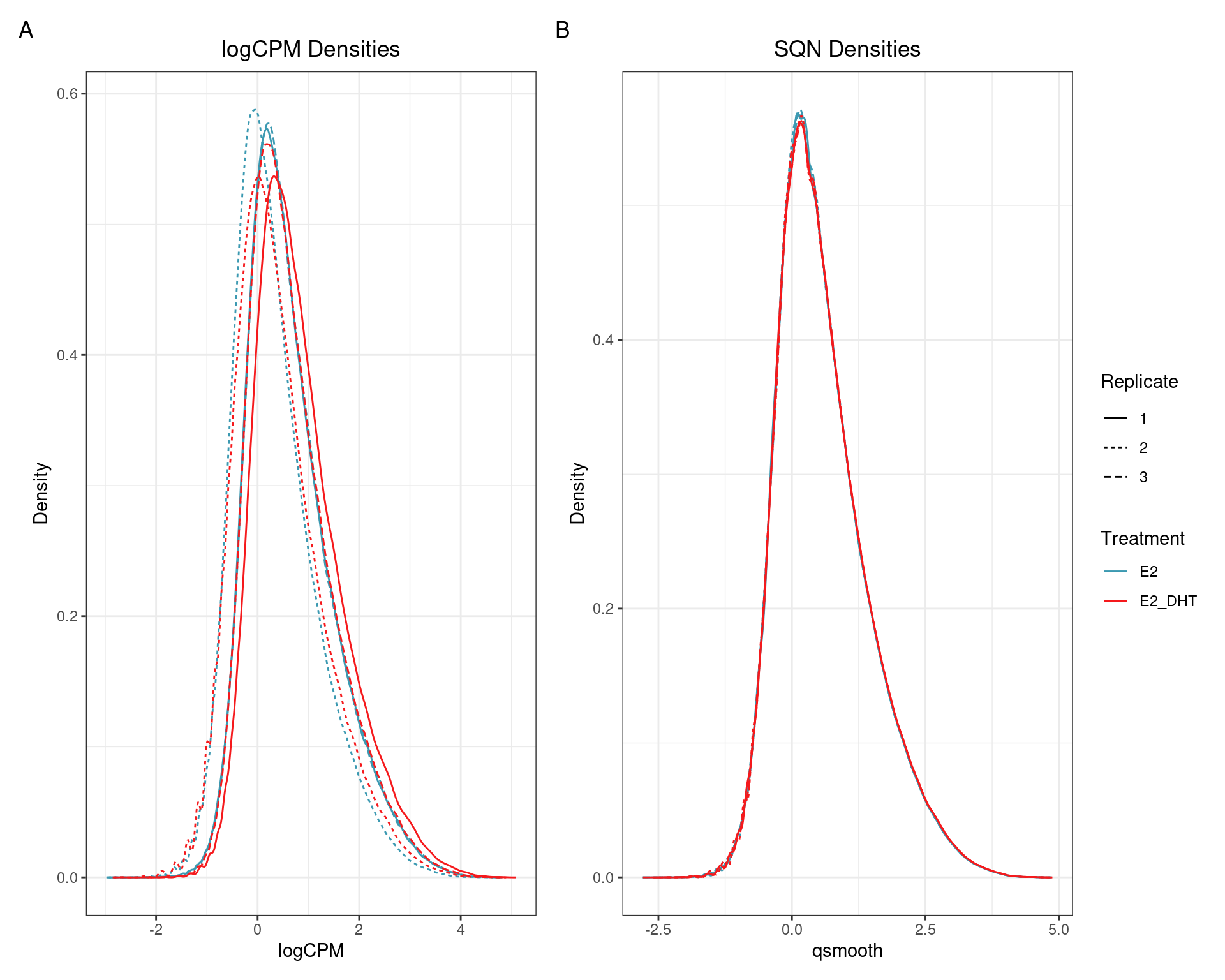
(quantro: \(p_{med} = 0.65\); \(p_{dist} < 5e4\))
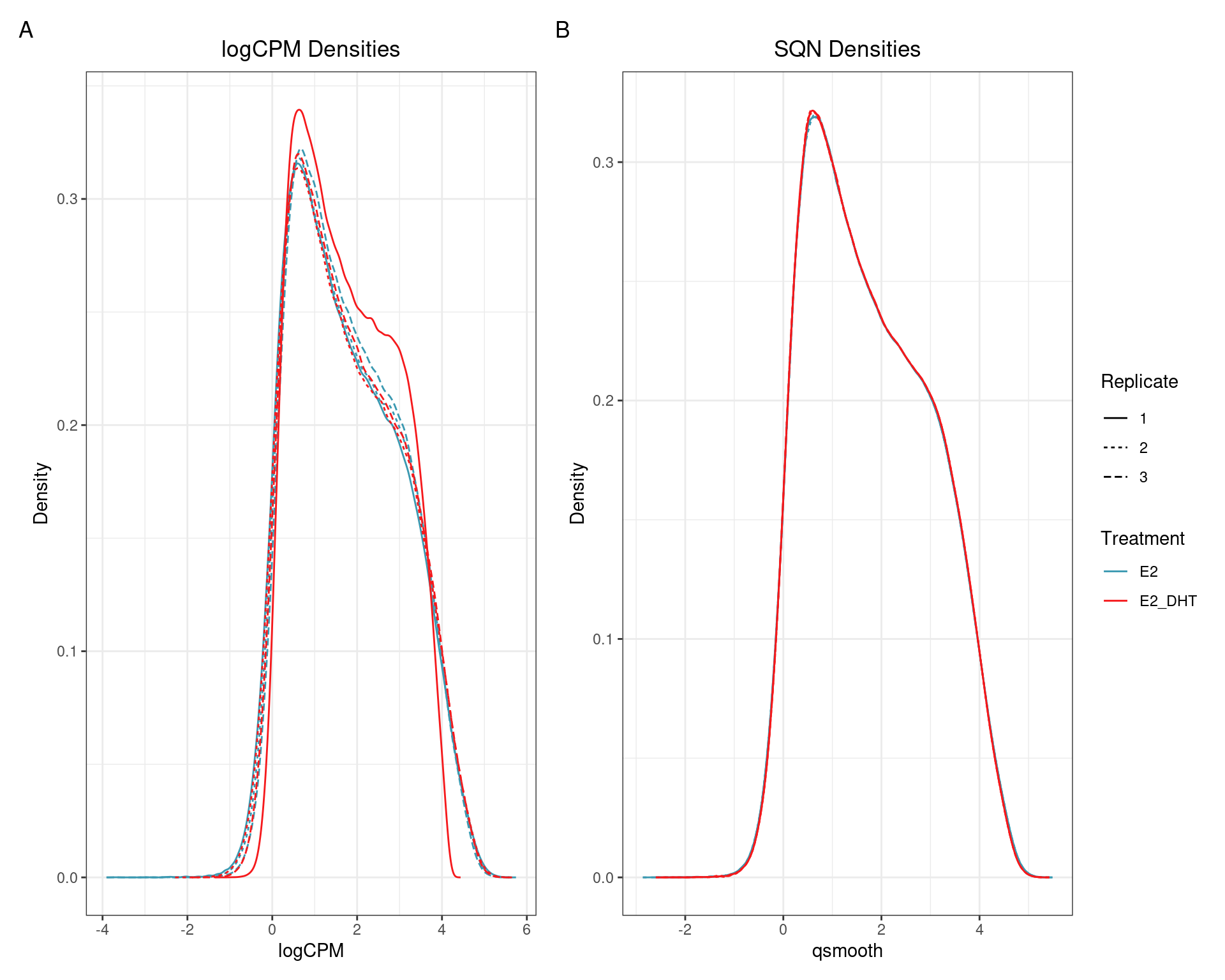
(quantro: \(p_{med} = 0.84\); \(p_{dist} = 0.49\))
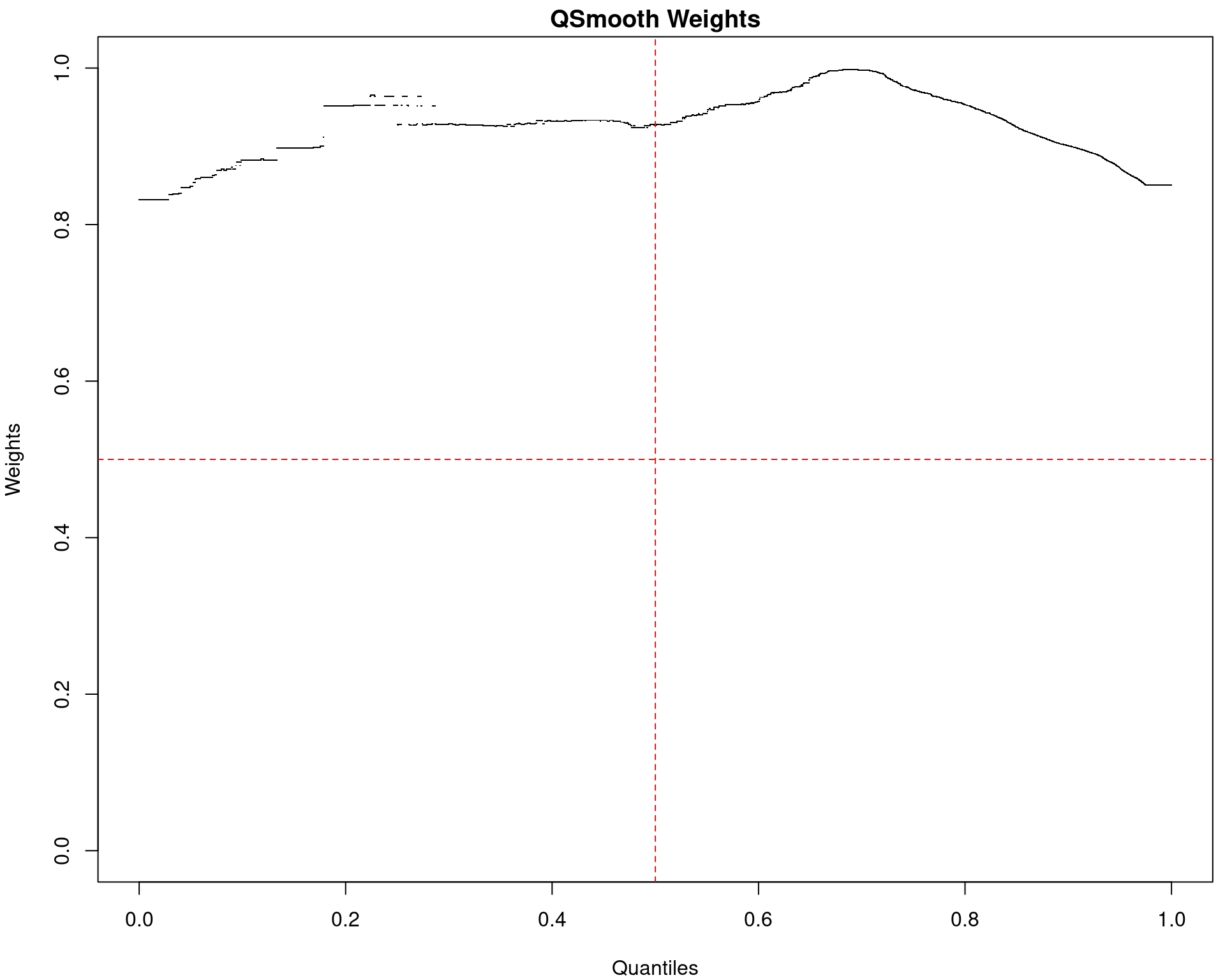
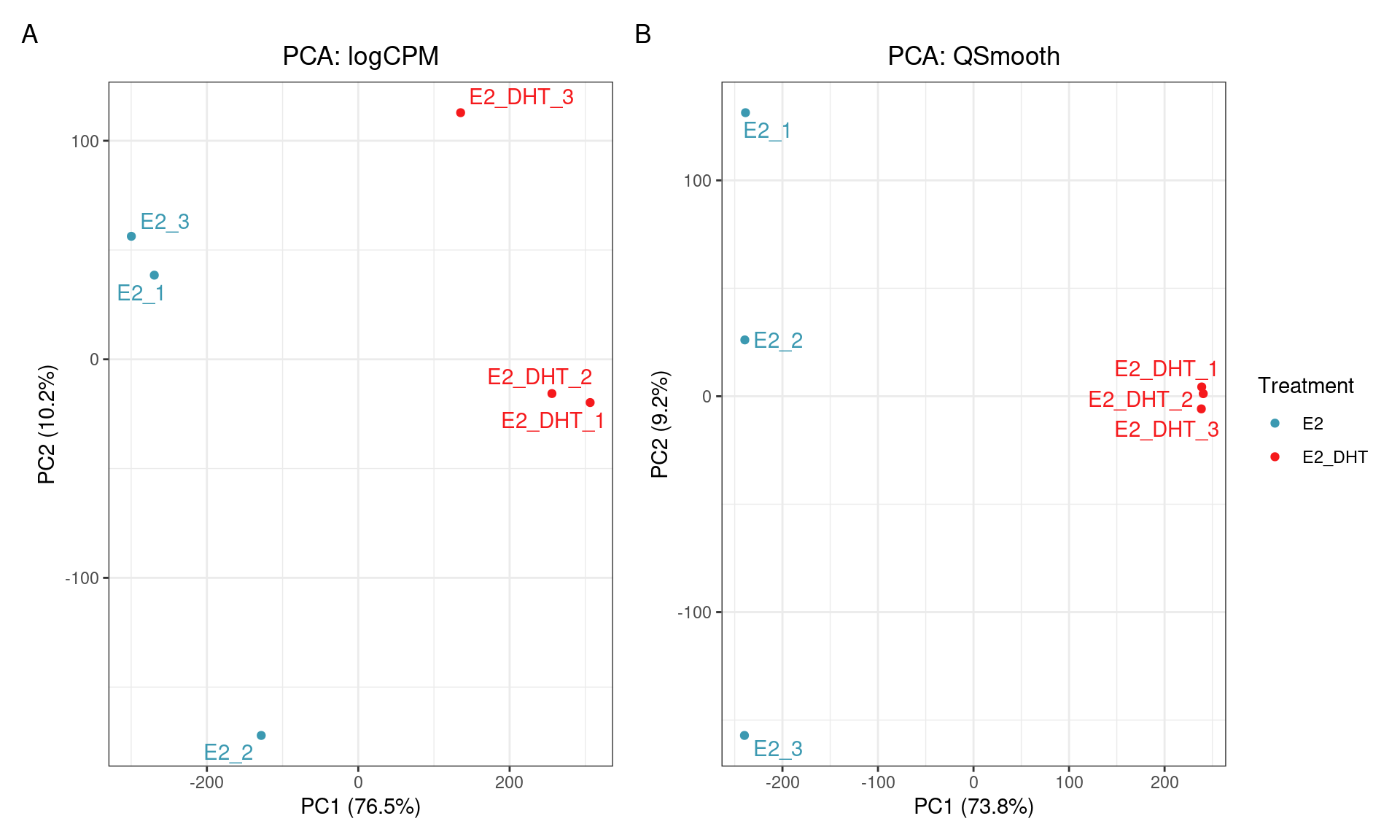
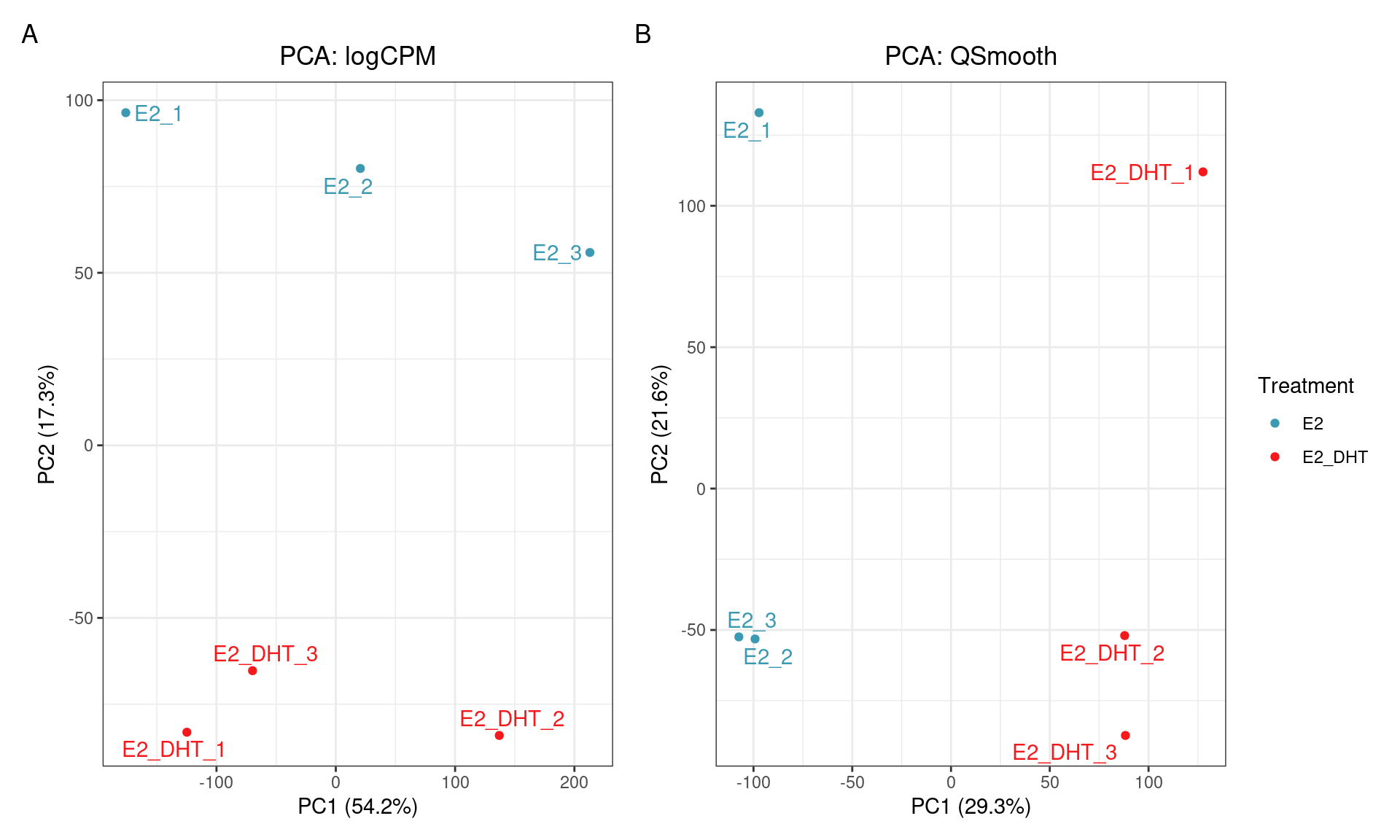
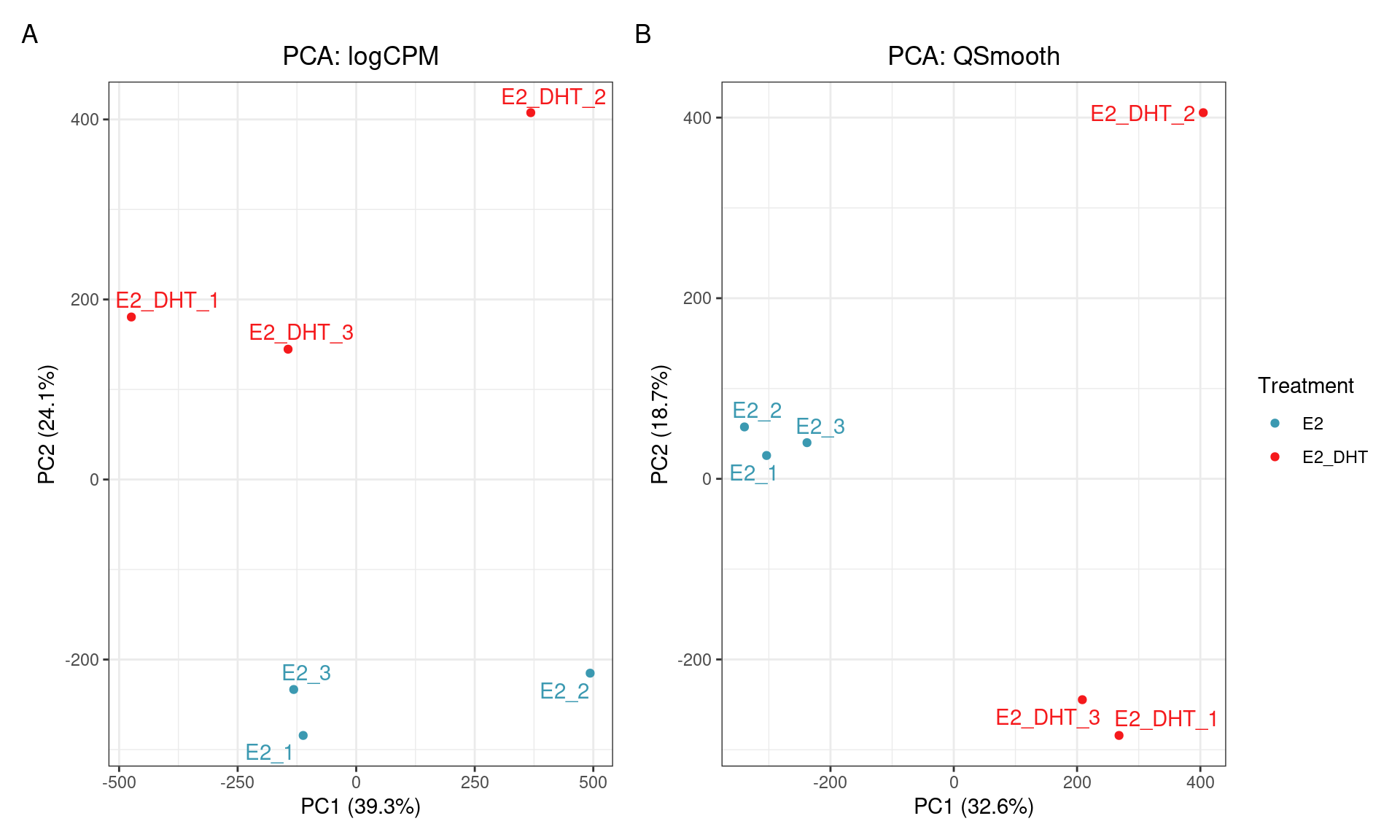
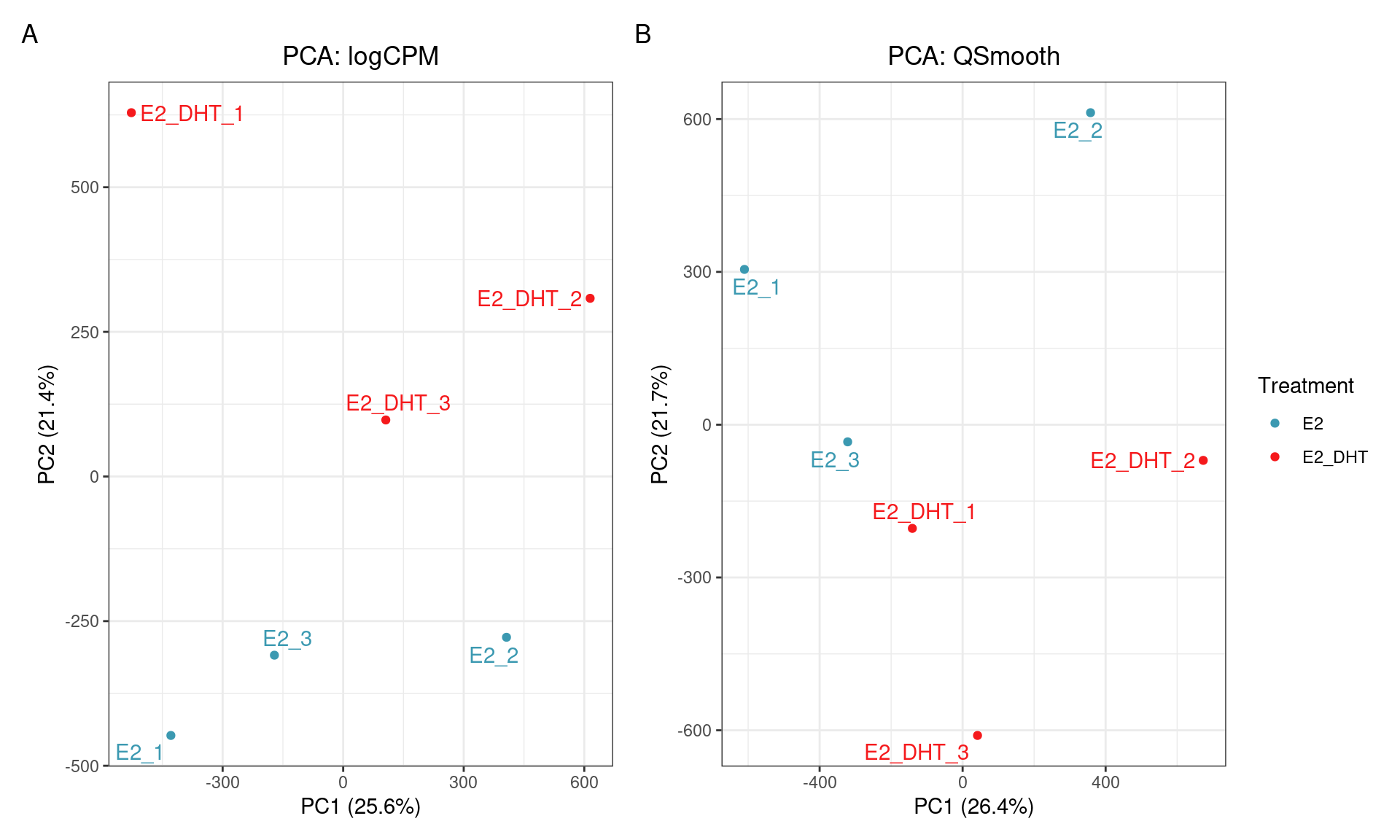
SQN: PCA
SQN
- Reducing replicate variability should yield more power
- Should be a clear improvement over library size alone?
limma-trendsimilar tovoom- Uses logCPM
- Manages mean-variance relationship
- SQN appears viable even when no real difference between groups
- Doesn’t seem to introduce any artefacts
- Could TMM also be applied within groups?
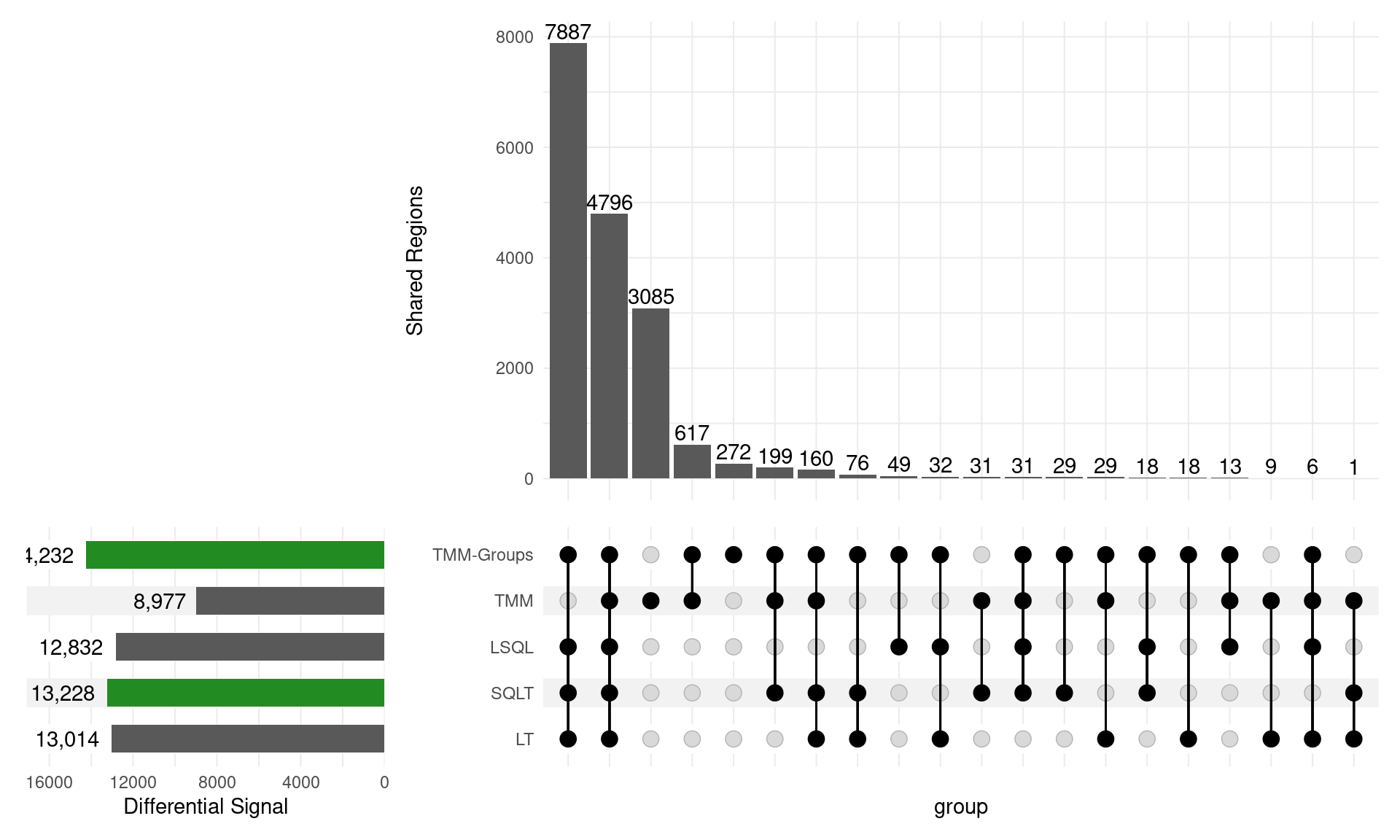
Method Comparison
Performed a comparison using
- LSQL: Library Size \(\rightarrow\)
edgeR::glmQLF() - TMM: (across all samples) \(\rightarrow\)
edgeR::glmQLF() - TMM-Groups: (within groups) \(\rightarrow\)
edgeR::glmQLF()
- LT: logCPM \(\implies\)
limma-trend - SQLT: qsmooth \(\rightarrow\)
limma-trend
Performed using
- \(H_0\) logFC = 0
- \(H_0\) |logFC| < \(\log_2 1.2\) (McCarthy and Smyth 2009)
All merged results using harmonic mean
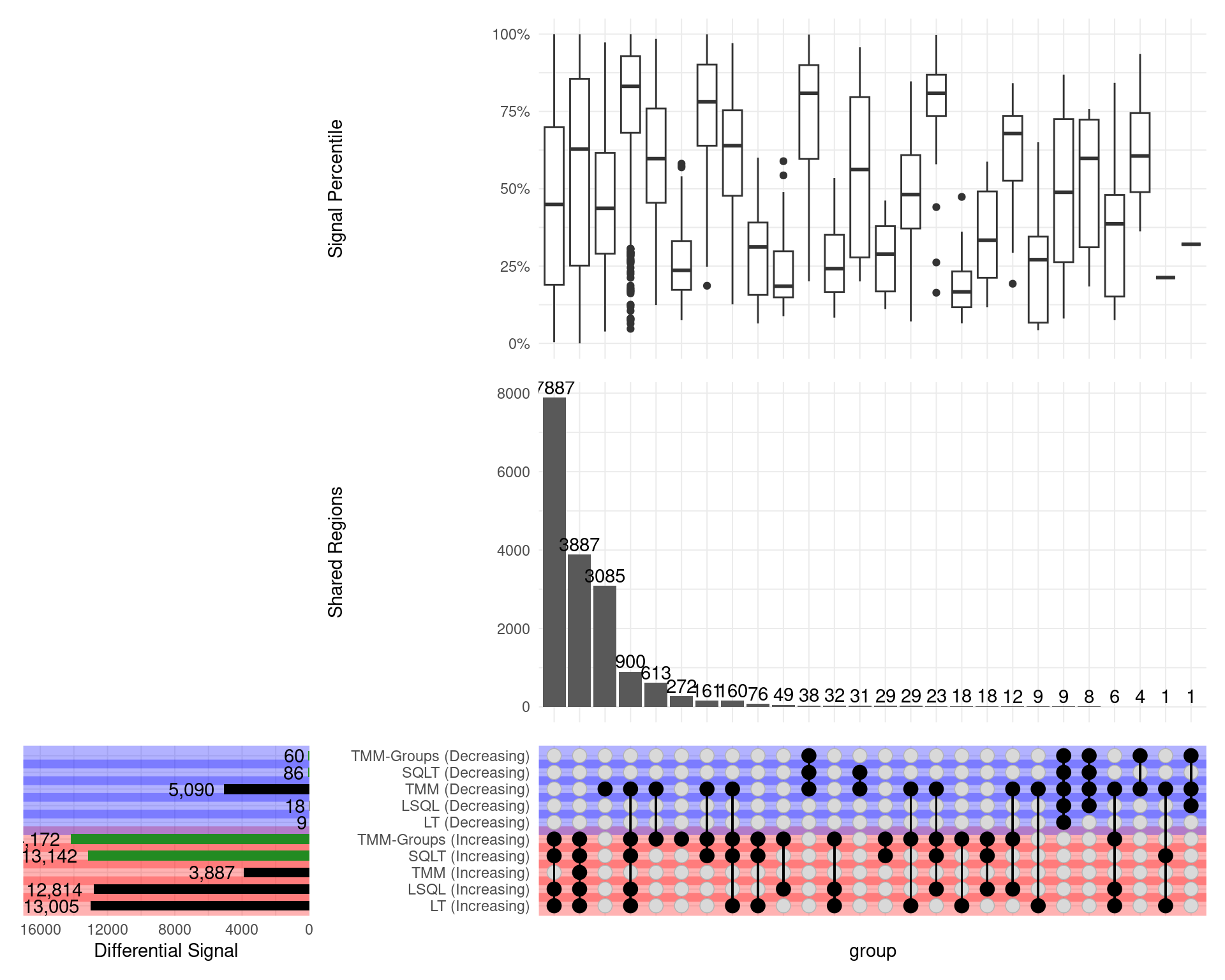
Overall Results
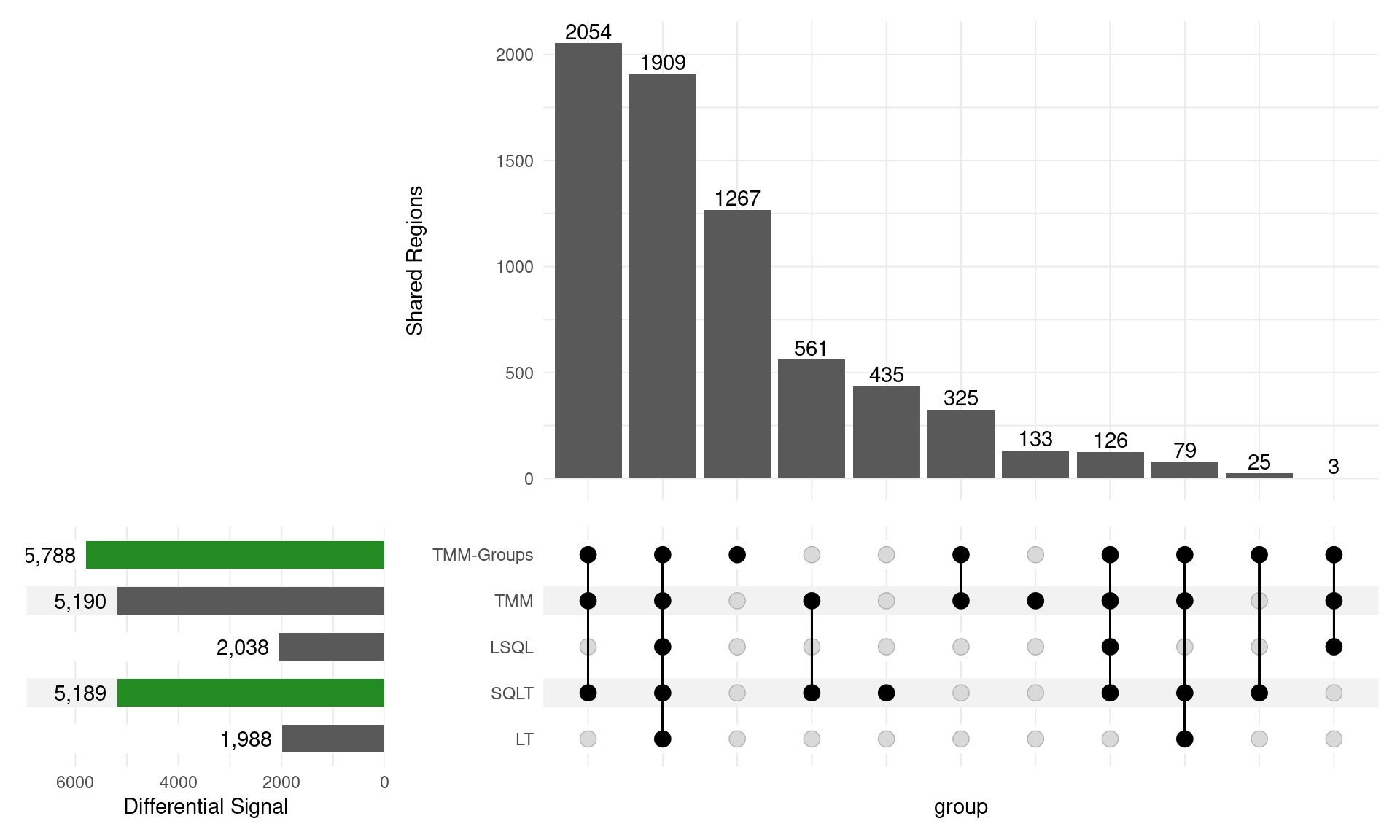
Directional Results
All Good So Far
- Group-based strategies looked best where expected
- SQLT appears poor for H3K27ac (??)
- How do the methods compare when using
treat(McCarthy and Smyth 2009)- \(H_0\) |logFC| < \(\log_2 1.2\)
- Known to control the FDR better than post-hoc filtering using logFC
- There was a drastic difference
- SQLT lost significant power compared to TMM-groups
- Had been competitive so far
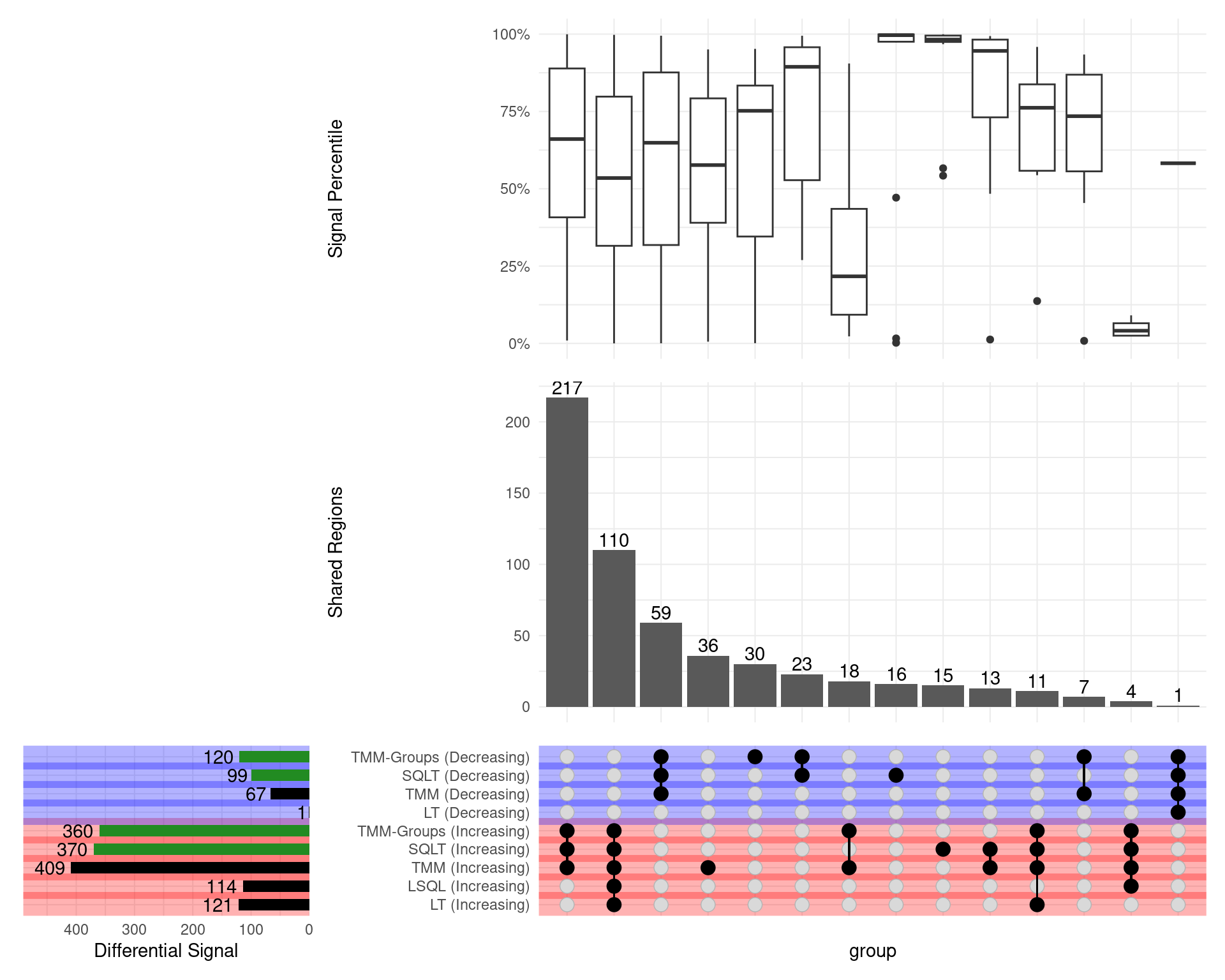
GATA3
\(H_0\): logFC = 0
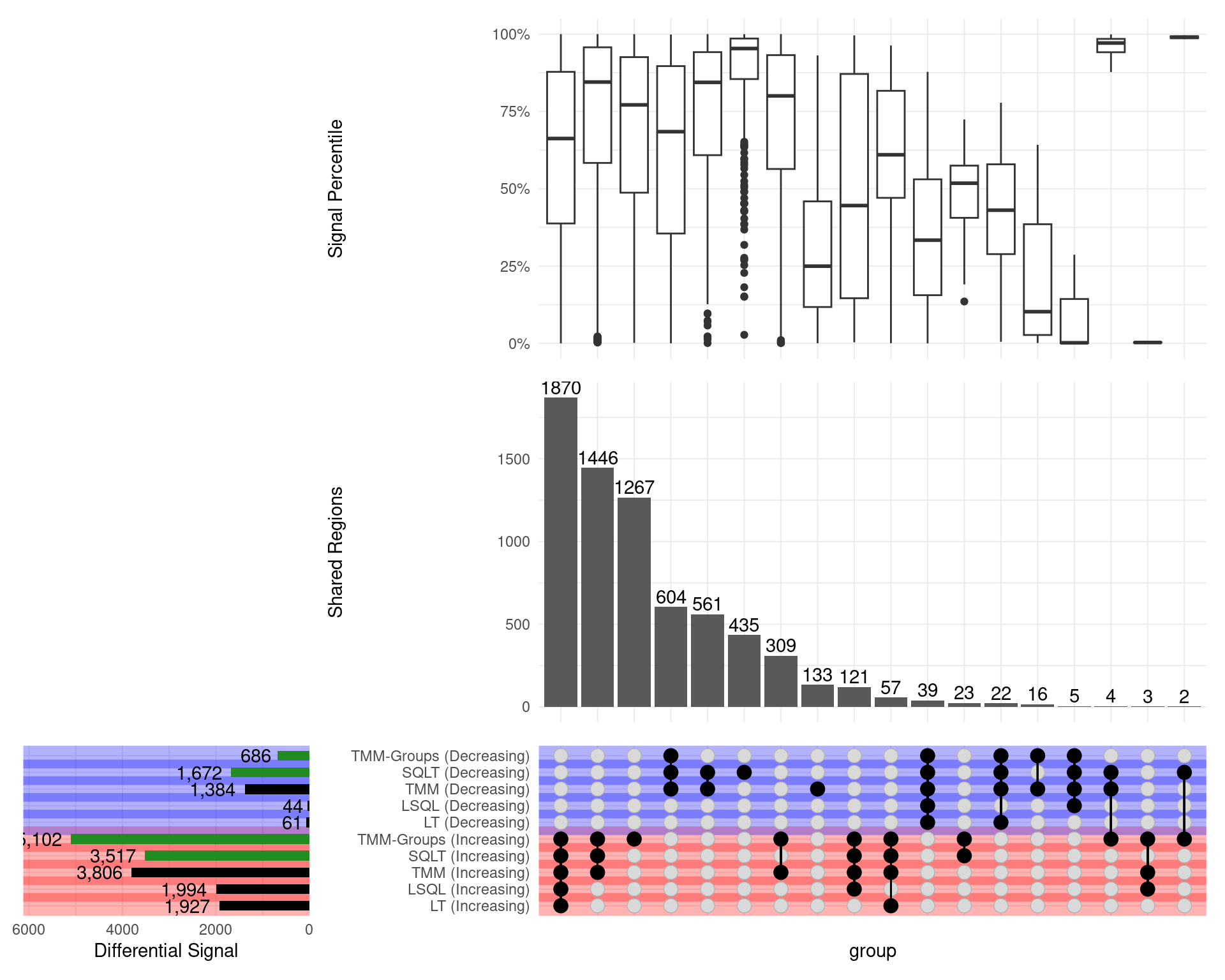
- SQLT has the most decreasing sites
- Less competitive for increasing
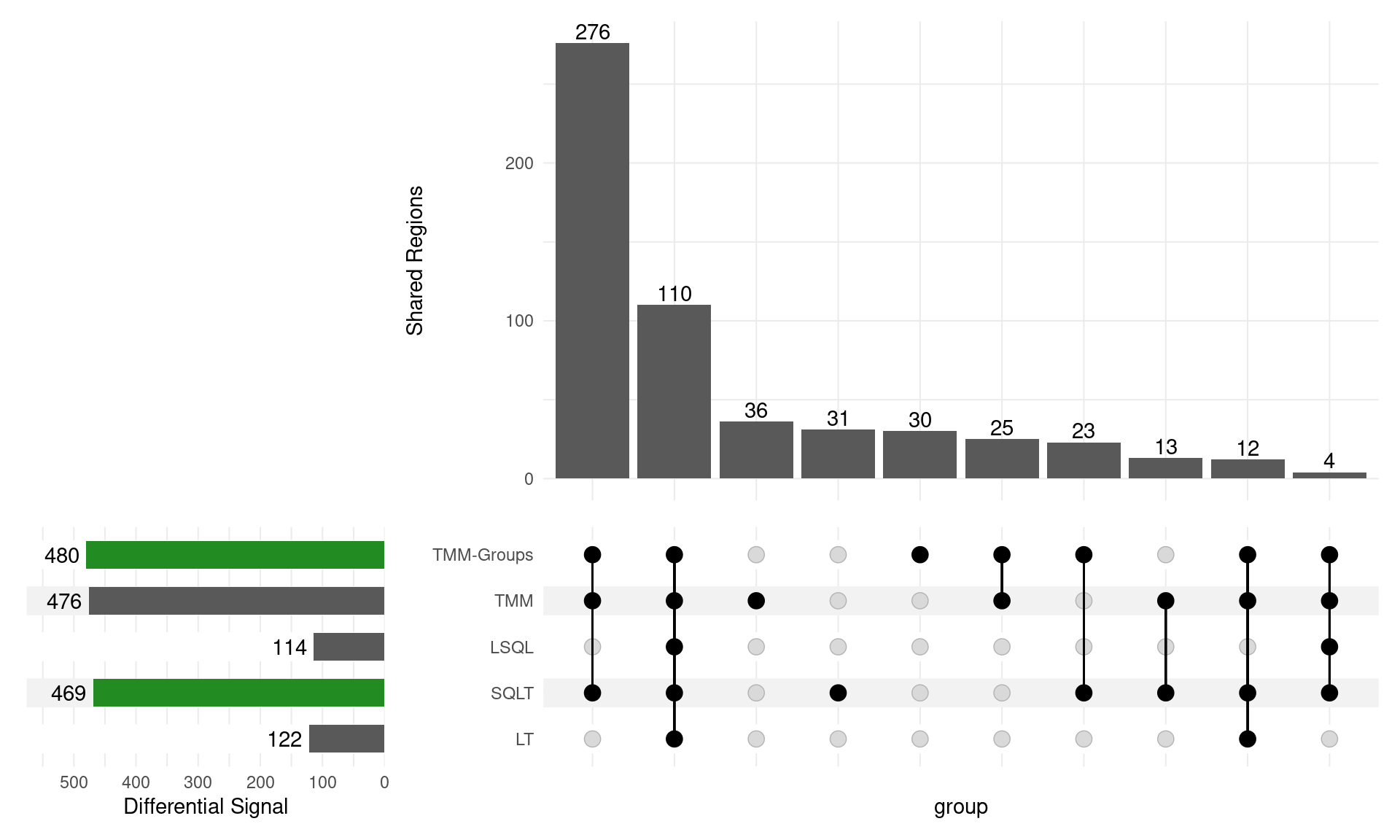
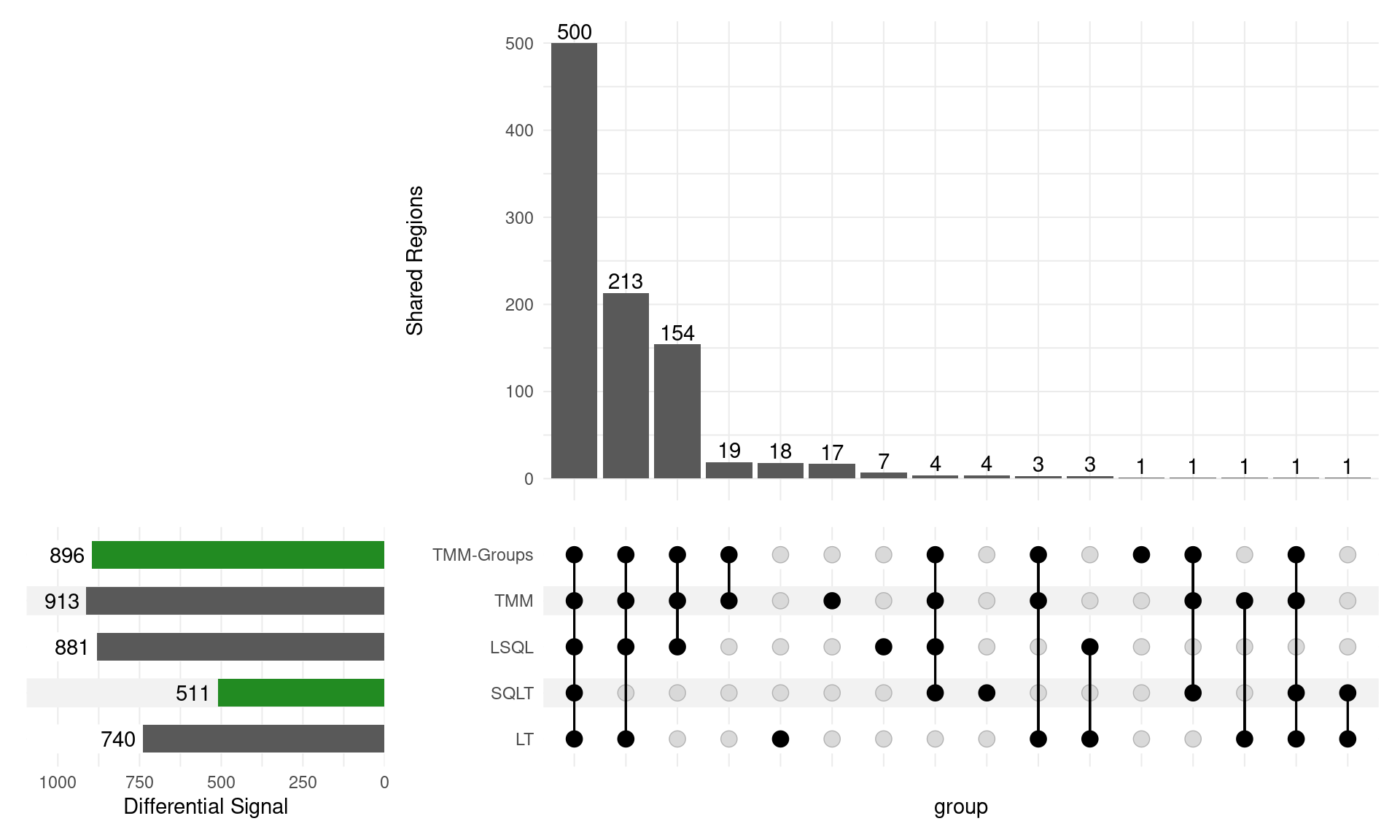
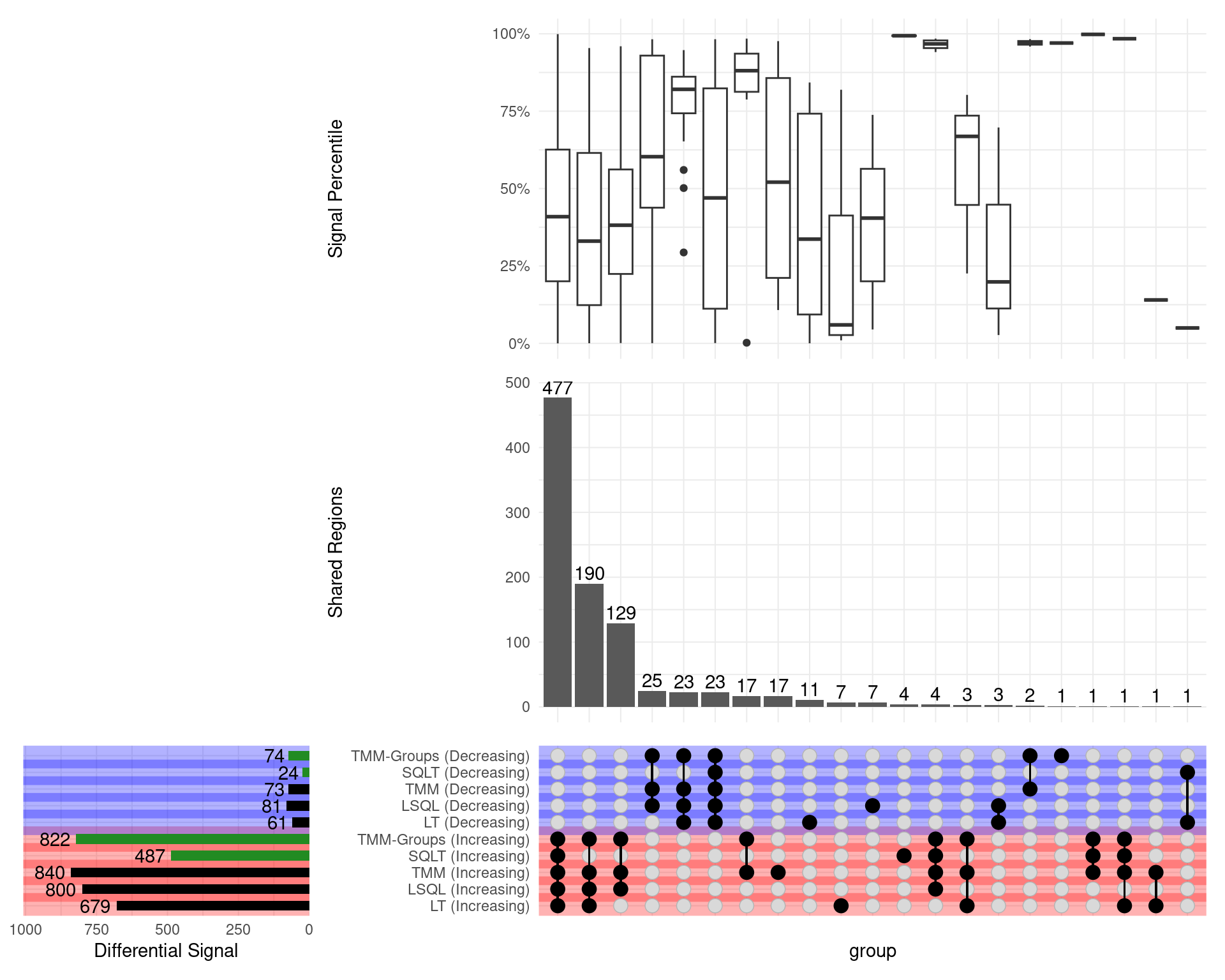
GATA3
Range-based \(H_0\)
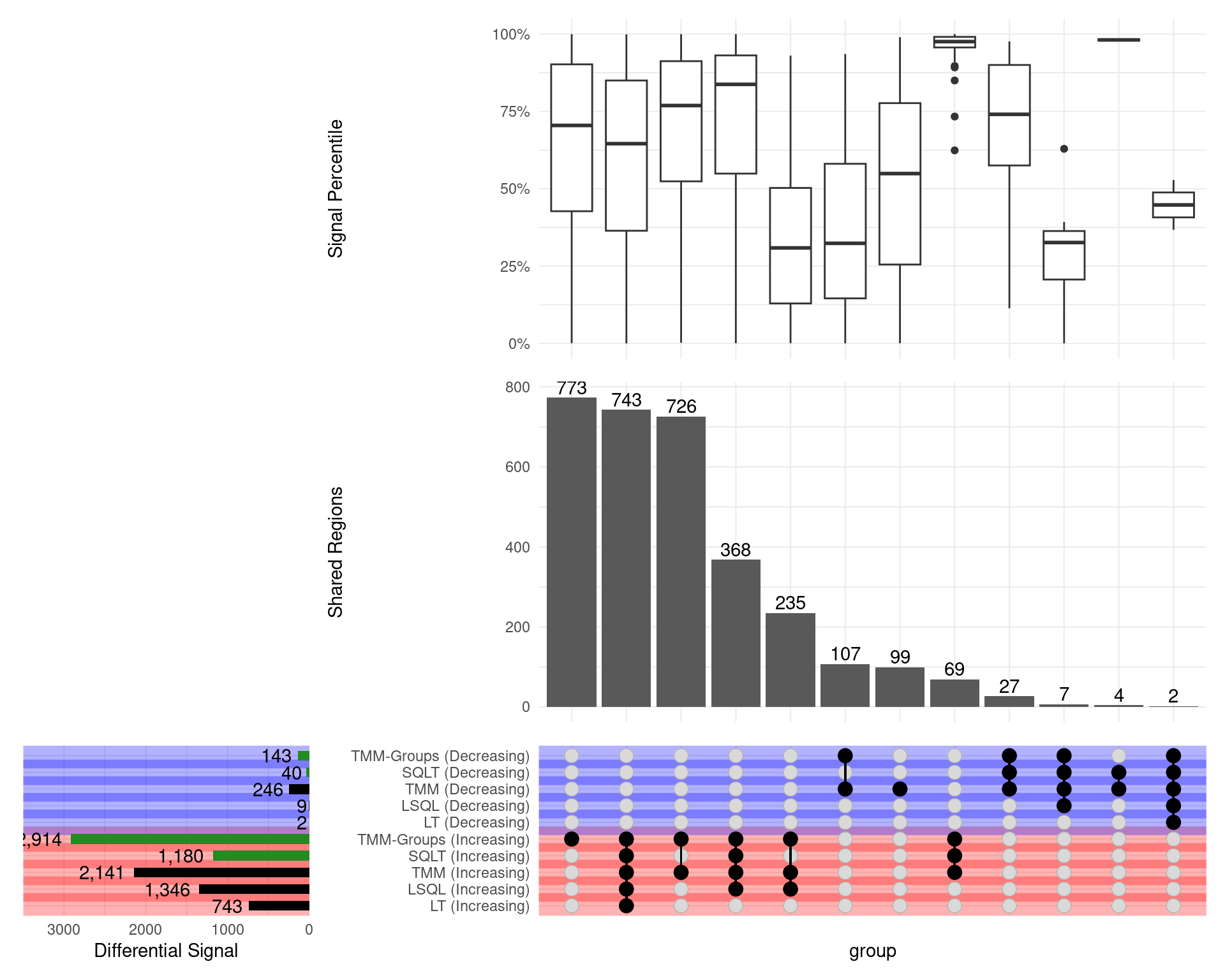
- SQLT now has only 40 down sites (from 1672)
- TMM Groups only dropped from 686 to 143
MA Plots: GATA3
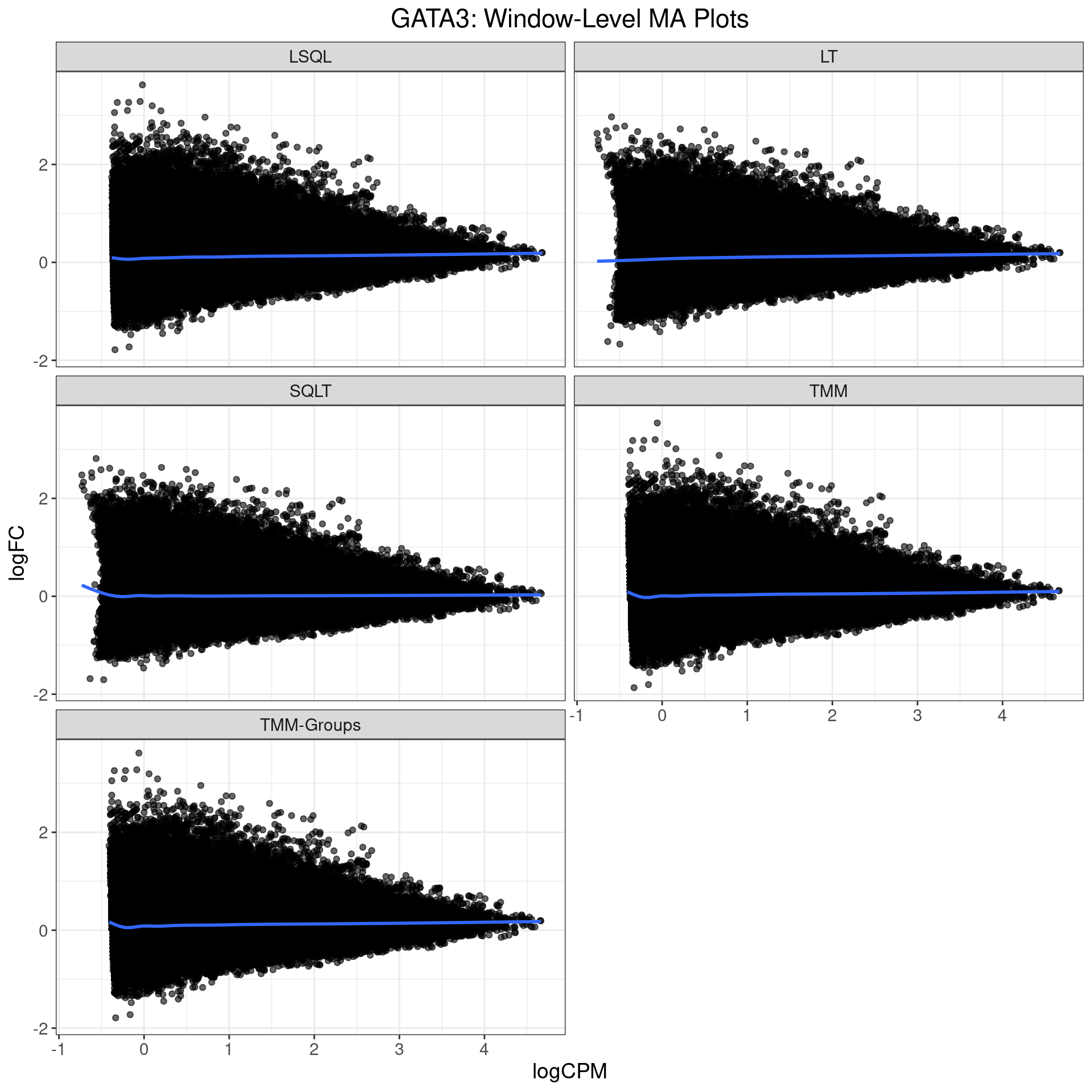
- Small positive bias in Library Size only (LSQL & LT)
- Small positive bias in TMM & TMM-Groups
- Is this why more gained sites?
- No apparent bias in SQLT
- Is ‘under-performance’ actually good?
- SQLT also appears more range-restricted
MA Plots: H3K27ac
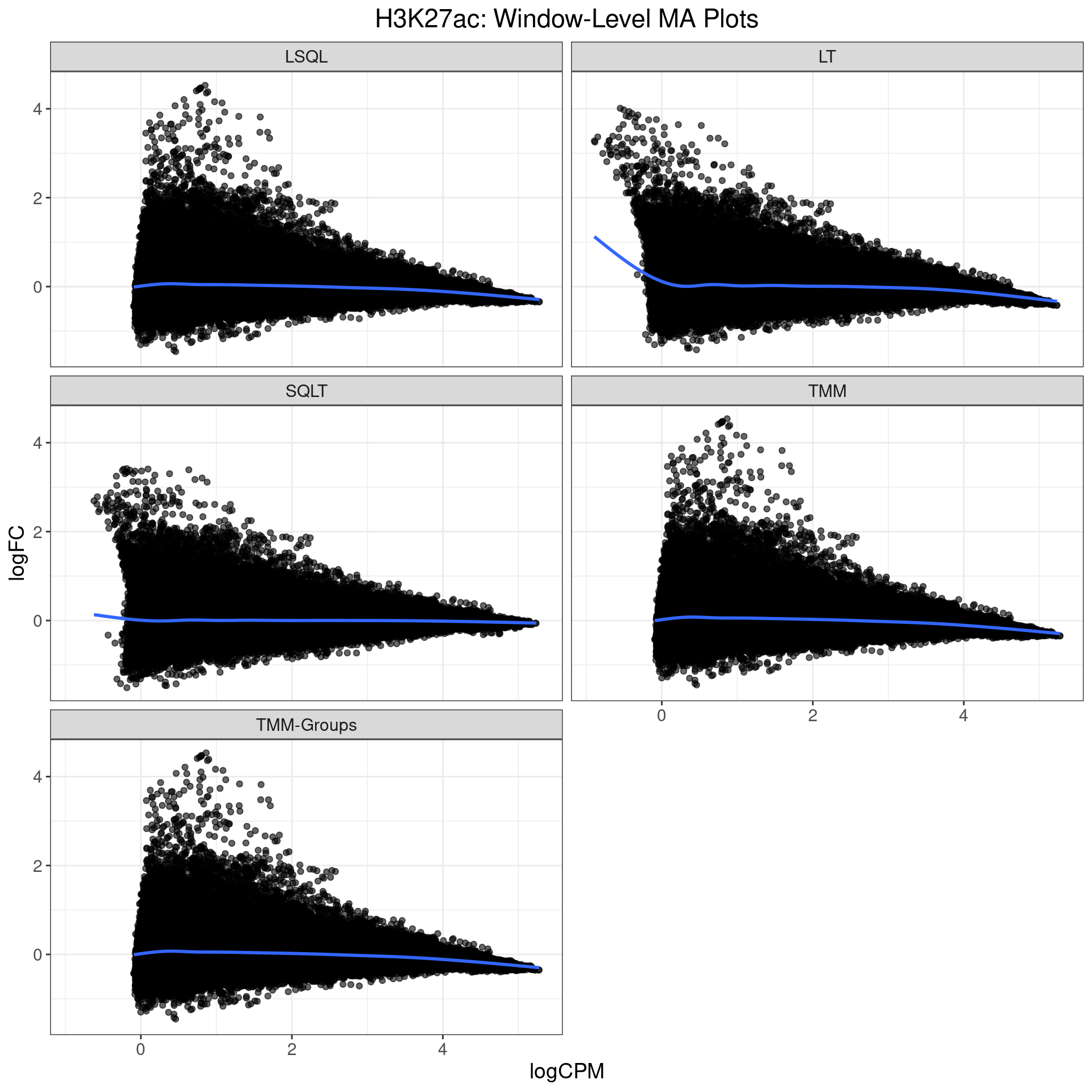
- Bias in all methods bar SQLT
- Most likely due to outlier sample
- Doesn’t quite explain the results
- Difference was mainly in Gains
logFC Comparison: LSQL Vs LT
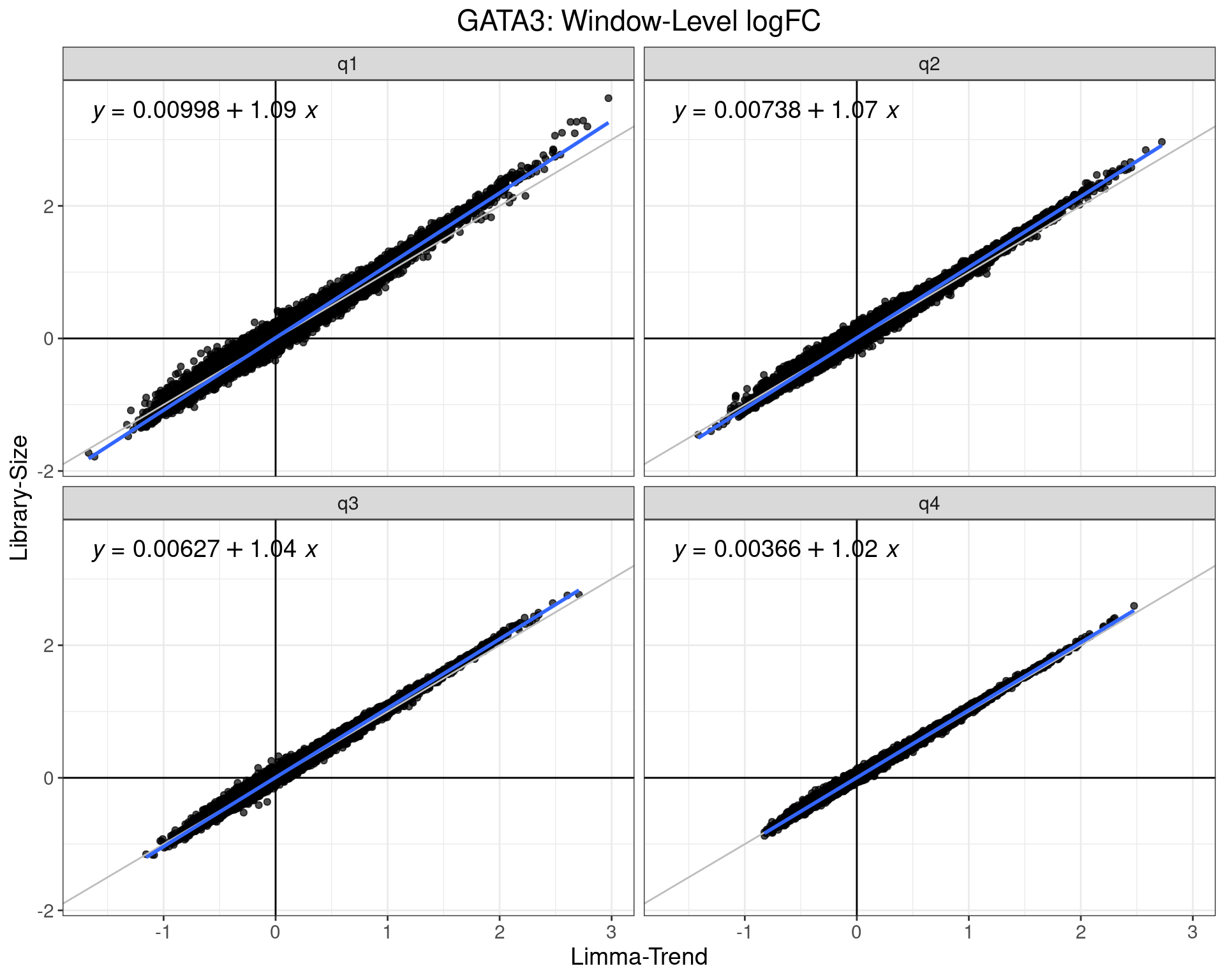
- Estimates from GLM-QL model are consistently higher
- Far more so in the lowest signal quartile
- Similar patterns in most datasets
logFC Comparison: SQLT Vs TMM-Groups
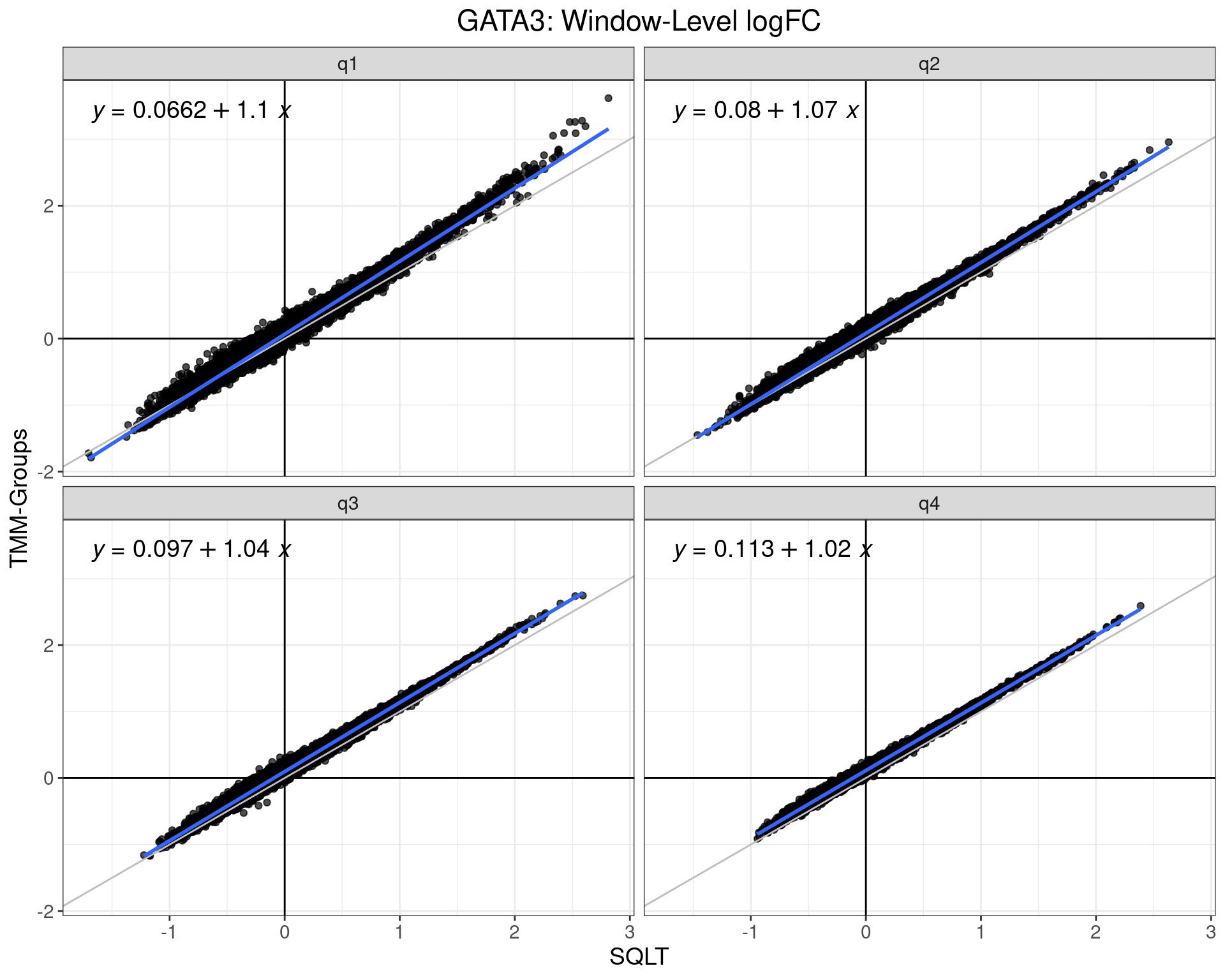
- Effect is even more pronounced for estimates using within-group normalisation
- Do I need to change the
treatthreshold?- Does GLM-20% = SQLT-18% in high quartile
- Does GLM-20% = SQLT-10% in the low quartile?
Final Thoughts
- Not all estimates of logFC are the same
- Becomes important when using
treat - 20% in GLM-QL \(\neq\) 20% under
voomorlimma-trend
- Becomes important when using
- GLM-QL approaches will give more significance at the low end
- Is that what we really want?
- May well be the same for RNA-Seq
- Is it the
prior.countsrestricting the range for logCPM?
- Is it the
Final Thoughts
- Within group normalisation was effective (quantro \(p < \alpha\))
- Within groups also helped for ‘different distribution’ datasets
- All-Sample TMM normalisation otherwise?
- Some bias was evident at the window level
- SQLT might still be viable
- Didn’t appear to disadvantage if not required
- Reduced technical variability nicely (PCA)
- Less bias at the window-level
- More likely to capture higher-signal results
- Fewer sites \(\implies\) but is this good?
Final Thoughts
- Harmonic Mean \(p\) still looks really good
(Did I mention my 100x speed-up?)
References